A COVID-19 Infection with Positive Sputum and Negative Nasopharyngeal/Oropharyngeal Rapid Antigen-Based Testing: A Case Report and Pilot Study
Abstract
Current covid-19 antigen testing is primarily carried out by obtaining a
specimen via nasopharyngeal/oropharyngeal swab and performing a rapid lateral
flow immunoassay (lfia) disposable micro-device or related immunoassays. In
this case report and pilot study, we propose the application of a modified
pretreatment antigen-based method for sputum, using at least one detergent and
one digestive enzyme. This method exposes hidden or masked antigenic sites of
viral specimens or lingering fragments of viral proteins present in a complex
biological matrix using a home-based rapid immunoassay for covid-19. The
modified technique can enhance detection sensitivity of lfia by making visible
the resultant test band, from viral-containing sputum samples, that would otherwise
not be seen. This protocol thereby overcomes a false negative result which may
occur when using a nasopharyngeal/oropharyngeal swab specimen from a patient
with mild symptoms of covid-19 and/or low viral load. This pilot study
pretreatment extraction sputum procedure is simple, non-invasive, rapid,
inexpensive, accurate, and may provide increased sensitivity, as well as
specificity in the detection of covid-19 antigens for several weeks or even
months.
Keywords: covid-19; sars-cov-2 virus; respiratory disease biomarkers; point-of-care
testing; lateral flow immunoassays; sputum proteomics.
Introduction
In the last decade, there has been a marked
improvement in the availability of laboratory and point-of-care tests for the
diagnosis of respiratory virus infections1.
Diagnostic tests for respiratory viral infections traditionally use nasopharyngeal
and oropharyngeal samples2,3. In the case of diagnosing coronavirus
disease 2019 (covid-19), the most common methods are the detection of severe
acute respiratory syndrome coronavirus 2 (sars-cov-2) antigen-based lfia and
rna assays in nasal and throat swab specimens. These techniques use a sampling
protocol that can be stressful, especially in children and when repeated
testing is required4. Unfortunately, these procedures have a rate
of false-negative results that might enable convalescent covid-19 patients to
meet the criteria for discharge from hospital and/or release from quarantine,
resulting in the spread of disease5,6.
Sputum is rarely used for viral testing7
given its viscous nature, making it difficult to process with automated
clinical microbiology laboratory equipment8. A type
of mucus secreted in the lower airways of the respiratory tract, known as
phlegm or sputum, is a selective barrier to particles and molecules, preventing
penetration to the epithelial surface of mucosal tissues. In general, mucus is
a complex hydrogel barrier located in the airways, gastrointestinal tract,
reproductive tract, and eyes9. The mucus is continuously produced,
secreted, and finally digested, recycled, or discarded; its main functions are
lubrication of the epithelia, maintenance of a hydrated layer, exchange of
gases and nutrients with the underlying epithelium, in addition to being a
protective barrier against pathogens and foreign substances9.
A recent study reported that thick, gummy
respiratory secretions are at the heart of severe covid-19 and are comparable
to the notoriously thick and tenacious sputum produced by patients with cystic
fibrosis10, calling for a more serious investigation
into the diagnostic value of sputum specimens. In fact, among the biological
matrices that are informative of lung status, sputum has gained growing
interest over the past decade for the investigation of several pulmonary
disorders11. Healthy mucus contains 3% solids, while
mucin hypersecretion or dysregulation of surface liquid volume may increase the
concentration of solids up to 15%, resulting in viscous and elastic mucus that
is not cleared. Mucus dysfunction occurs in virtually all inflammatory airway
diseases12.
With the advent of molecular methods, it has
been found that by processing sputum, sputum testing information adds
approximately 11% to the diagnostic yield for the detection of many common
respiratory viruses8. In particular, the use of sputum samples for
the detection of respiratory viruses has been helpful in patients with chronic
obstructive pulmonary disease (copd), asthma, and cystic fibrosis (cf), as well
as other pulmonary disorders. Current evidence suggests that sputum processing
may even be required for diagnosis, given that certain viral pathogens such as
h1n1 influenza, sars-cov-2 coronavirus, and mers coronavirus typically present
in the lower respiratory tract may be absent in upper airway secretions or
nasopharyngeal samples8. A recent study in india is even recommending
the use of sputum testing as the new mass screening method for individuals
affected by covid-193. Similarly, because sputum is more reliable
and has a lower false-negative rate than throat swabs, studies in china are
recommending the use of induced sputum as a test specimen, because it is more
accurate for the confirmation of covid-19 and is safer as a criterion for releasing
patients from quarantines13. It is evident that studies on sputum content
have helped improve understanding of chronic airways disease as it can identify
the presence and type of microorganism, which can indicate the severity of
airways disease and thereby aid treatment and management options14,15.
Since sputum has a complex, compact molecular
polymeric hydrogel structure, it is desirable to develop a simple method to
forge better accessibility to its internal content that may be sheltered,
hidden, or masked within the mucus network barrier: antigenic viral proteins or
other viral constituents, pathogenic microorganisms, or toxic materials.
Manipulation of this polymeric structure is the key to widespread sputum use in
the detection of viral proteins by lateral flow immunoassay test and other
immunoassays. Previous studies have reported that the sputum pretreatment
process presently used in the laboratory is a significant obstacle to on-site
diagnosis of respiratory infections. The current pretreatment procedure is
complex and labor intensive16.
As such, this paper describes a pilot study of
a sample pretreatment extraction procedure applied in conjunction with a
covid-19 lateral flow immunoassay test allowing the release of significant
sputum content, therefore yielding a more accurate test result. The simplicity
of the assay is based on the use of at least one detergent and no less than one
digestive enzyme to disrupt/lyse the sputum matrix and assay of the released
components of interest using a lateral flow immunoassay platform. This improved
sample preparation procedure can also be employed for other antigen-based lfia
or related immunoassay screening tests. This method allows for the simultaneous
detection of respiratory viruses and other pathogens (bacteria, fungi),
together with sputum biomarkers, in a single and highly multiplexed broad
assay.
Detailed case description
The rationale for the research described in this paper was based on the experience of a 75-year-old male patient looking for answers to certain symptoms that appeared to be covid-19. The symptoms included fatigue, minor headache, and productive cough without fever, rhinorrhea, loss of taste or smell, nausea, or vomiting. Physical examination by an emergency medical doctor at an urgent care clinic (brunswick urgent care, east brunswick, new jersey, u.s.a.) and an on-site antigen-based rapid lfia test were performed at the same clinic, using nasopharyngeal/oropharyngeal (np/op) swab specimens; confirmatory rt-pcr testing was also sent to a specialized clinical laboratory (quest diagnostics-healthcare company, east brunswick, new jersey, u.s.a.). Additional pcr testing was performed at another private laboratory (ez clinical laboratory, east brunswick, new jersey, u.s.a.). The results of both the rapid antigen-based test and the rt-pcr test performed on the same day on np/op collected samples were reported negative. The additional rt-pcr test performed at another private laboratory was reported invalid, on sputum specimens that were kept in a freezer in the presence of a detergent until the day of testing.
Conventional protocol applied to repeat the antigen-based test
When the patient experienced persistent symptoms, more studies were decided to be performed. Experiments were carried out using commercially available covid-19 at-home antigen-based rapid lfia diagnostic tests specifically designed for the detection of sars-cov-2 nucleocapsid protein antigen n, one of the most produced proteins of the sars-cov-2 virus. The kits were obtained from three different suppliers: flowflex covid-19 antigen home test from acon laboratories, inc., san diego, california, u.s.a.; ihealth covid-19 antigen rapid test from ihealth labs, inc., sunnyvale, california, u.s.a.; and sars-cov-2 rapid antigen test from roche diagnostics, indianapolis, indiana, u.s.a. experiments testing the various samples were performed using all three kits, and the results were comparable and reproducible. Nasopharyngeal swab samples were collected in our laboratory from the patient. The lfia procedure for detecting the presence of the coronavirus was performed as indicated by the kit manufacturers, using disposable sterile nasal swabs included in the kit provided by the suppliers. The swab was introduced into the nostril as instructed by the manufacturer protocol, followed by the insertion of the swab into a tube containing an extraction buffer carrying a surfactant. After the appropriate extraction procedure and mixing of the sample material absorbed and retained onto the swab, three to four drops of the solution were removed from the squeezable plastic tube and gently applied to the sample well of the platform or cassette of the manufacturer’s test kits, known as the sample pad17. The entire procedure to perform the lfia tests17-20 were carried out in our laboratory at a temperature of approximately 25- degrees celsius. The process begins with a lateral flow chromatographic migration that occurs for the applied sample onto the sample pad containing sars-cov-2 antigens. The sample viral antigens then bind to the matching antibodies present in the corresponding area of the platform, known as the conjugated pad. These antibodies are labeled with gold nanoparticles (aunp). As migration occurs by capillary action, viral and non-viral constituents of the processed-extracted sample flow progressively through the strip or platform from the sample pad to the conjugated pad, passing through the membrane pad or detection zone, and ending in the absorbent pad needed to maintain the movement of fluid. The role of the absorbent pad is to wick the excess reagents and to prevent backflow of the liquid. The final antigen-antibody reaction occurs at the membrane pad or detection zone: a porous membrane (usually composed of nitrocellulose). This membrane contains immobilized antibodies in the t lines allowing the binding of the sample analyte (target antigen), already bound to another labeled antibody pre-embedded at the conjugated pad, with the corresponding immobilized antibody. This double-antibody sandwich-like method interaction results in the formation of a visible color17-20. The read-out, represented by the colored lines with different intensities, can be assessed by eye or using a dedicated reader. Depending on their size, shape, degree of aggregation, and local environment, gold nanoparticles can appear red, blue, or another color. Gold nanoparticles (aunp) are used as color markers with unique optical properties, extraordinary chemical stability, robustness, and high binding capacity for biomolecules21. The control (c) line should be visible independently of the test result. When the test (t) line is visible (positive results), it indicates the presence of viral antigens in the specimen and implies that the person tested is in fact considered to be infected by the sars-cov-2 virus. If no color is observed (negative results), then the person is considered not to be infected by the virus.
Other specimens, such as oropharyngeal swab sample, buccal swab sample, saline mouth rinse-gargle, and saliva were also tested, and the protocol was carried out identically to the nasopharyngeal swab specimen previously described using the conventional lfia as recommended by the manufacturer’s test kits. All reagents were sterilized before use. A negative result for the presence of sars-cov-2 virus was obtained when testing specimens obtained with a swab from nasopharyngeal, oropharyngeal, buccal, saline mouth rinse-gargle, and saliva samples. A strong visible colored band appeared in the control line, whereas no color was observed in the test line (figure not shown).
Alternative protocol applied to confirm diagnosis
Additional samples, such as saliva and sputum, were collected and a modified pilot study pretreatment extraction procedure was used to boost the release of antigenic viral particles from mucus fluid, and thus to facilitate the performance of the lfia tests. Collection of saliva and expectorated sputum was carried out in a sterile collector-reactor tube without using a swab. Collection of other samples (nasopharyngeal, oropharyngeal, buccal, and gargle) used as control to test the modified protocol, were obtained in the same way as described above using a swab specimen using the sterile components from the kit supplied by the manufacturers. However, the processing-extraction protocol was modified before performing the lfia test. The modified protocol consisted of adding at least one detergent followed by at least one proteolytic enzyme to the saliva and sputum samples to disperse and alter the mucus structure. The preferred detergent was triton x-100 (santa cruz biotechnology, inc., dallas, texas, u.s.a.) and the preferred digestion enzyme, a protease, was subtilisin a - alcalase® food grade (novozymes a/s, bagsvaerd, denmark). Alcalase is a versatile endoprotease providing very extensive hydrolysis. The modified extraction-lysis procedure developed for sputum and saliva was also applied to all other specimens tested.
The modified extraction detergent-protease sample preparation protocol was mainly designed to disrupt primarily the complex hydrogel, polymeric composition of a viscous sputum specimen. In the first week of symptom onset, the production of expectorated sputum was abundant and easily collected; thereafter, the amount of sputum produced declined steadily through the course of illness. Once the sputum was collected, triton x-100 and free-enzyme alcalase were added to the collection-reaction tube, and the mix was incubated at 25-degrees celsius for a period of 5 minutes to 2 hours. The tube was inverted a couple of times during this process using a constant and gentle rotation of the wrist. The tubes feature screw caps that provide a tight, secure seal. After the incubation period, the disrupted-extracted-lysed sample mix was allowed to settle by gravity or centrifugation to remove some formed debris. A portion of the supernatant was then placed in the sample pad of the lfia platform or strip to let the sample migrate to the absorbent pad.
Samples of saliva and sputum were collected in a sterile container early in the morning, before eating or drinking, and after rinsing the mouth with clear water for about 15 seconds to eliminate any contaminant in the oral cavity as described previously22. In the case of sputum, the same protocol was used, except that saliva was expelled first, the patient then breathed in deeply three times to cough at 2-minutes intervals until bringing up some sputum. The sputum was then released in a sterile, well-closed container obtained from a local pharmacy. About 1 ml of a triton x-100 solution was added to approximately 2 to 3 ml of saliva or sputum (v/v). The concentration of triton x-100 used ranged from 0.1% to 2.0% of total volume, of which 1.0% was the preferred concentration. The times of incubation at 25-degrees celsius of the mixed sputum-detergent-enzyme solution were determined as how the enzyme was used, either as a free solution enzyme or immobilized to a solid support and ranged from 5 minutes to 2 hours. Regarding the quantity of alcalase used, approximately 15-60 microliters of a free-solution enzyme were added to a total volume of 3 milliliters of collected sputum sample mixed in the detergent, making a ratio of about 0.5-2.0% of enzyme- sputum solution (v/v). Other ratios were also used, depending on the viscosity of the sputum sample. After incubation, a disrupted-lysed saliva or sputum solution (figure 1) was decanted or centrifuged followed by the addition of 3 to 4 drops of the supernatant to the sample pad of a lfia strip as previously described before. The lfia was then assayed for the presence of sars-cov-2 antigenic viral components. All other samples were collected separately and individually using a swab, which was then immersed into a tube containing a solution of a detergent and at least one proteolytic enzyme followed by mixing and incubation.
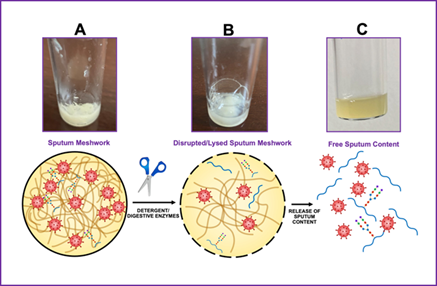
Figure 1. Diagrammatic representation of the sample preparation (disruption-extraction-digestion) procedure, whereby the nonionic detergent triton x-100 and the endoprotease alcalase are applied to a sputum specimen. As depicted in panel a, the sputum is a thick, rubbery, sticky, viscous, and gel-like meshwork. Sputum or mucus of the respiratory system contains numerous cells, cell debris, microorganisms, and chemical-biochemical entities. After adding the detergent and the protease, some disruption occurs (panel b) as influenced by time of incubation, quantity of the proteolytic enzyme, temperature, and ph of the solution. In most experiments, the temperature of incubation was 25 degrees celsius. This process resulted in a solution containing primarily soluble material and some precipitate of insoluble components (panel c). After decantation or centrifugation, the supernatant was tested for the presence of sars-cov-2 virus, or virus components, on a lfia platform or strip.
The results for all tested samples (nasopharyngeal, oropharyngeal, buccal, and gargle) using the modified protocol were negative (figure not shown), except for the sputum specimen that yielded positive for the presence of sars-cov-2 virus as shown in (figure 2). Panel a shows the results of a sputum sample incubated with the detergent and protease for a short time, usually 5 to 10 minutes. Panel b shows the results of a sputum sample incubated with detergent and protease for a longer time, usually 1 to 2 hours. The intensity of the band seen in panel a was weak; however, the band intensity of an aliquot of the same sample increased with a longer incubation time, as shown in panel b. This indicates that time, temperature, and enzyme concentration seem to be crucial for an optimal disruption-extraction-digestion of the sputum sample to be able to release the maximum content of the constituents trapped within the complex and difficult to disrupt meshwork barrier of the gel-like sputum.
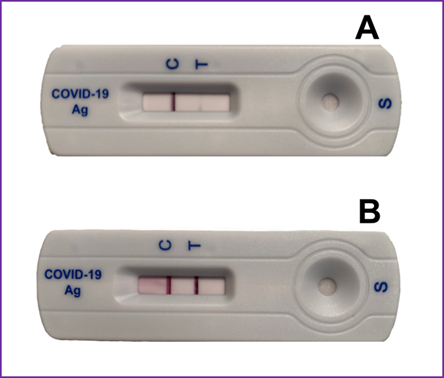
Figure 2. Diagrammatic representation of the sandwich format of a lfia test using the modified disruption- extraction sample preparation protocol for sputum. Panel a shows an aliquot of the sputum-detergent-protease mix tested at approximately 5 minutes of incubation at 25-degrees celsius. Panel b shows an aliquot of the sample mix incubated for about 1 hour at 25-degrees celsius.
Free-solution and immobilized alcalase
The endoprotease alcalase employed in the experiments reported in this paper was used as a free-solution enzyme or as an immobilized enzyme to a solid-support (figure 3). In most experiments, the amount of expectorated sputum was abundant in the first week of performing experiments and declined significantly after several weeks. However, there was always a small amount of sputum available for use in the experiments. The proportion of sputum to detergent was maintained to an approximately ratio of 2 to 3 parts of sputum to about 1 part of detergent (v/v). The covalent immobilization of alcalase on a solid support was carried out by previously described procedures for other enzymes or proteins23-25. Two methods were primarily used to immobilize alcalase to beads. The first method used an epoxy-activated beaded resin with a high density of epoxy-functionality (toyopearl af-epoxy-650m, tosoh bioscience llc, king of prussia, pennsylvania, u.s.a.). The procedure for immobilization via this method was carried out with a 50 mm sodium phosphate buffer, ph 7.0, with minor modifications of a method described elsewhere26. After immobilization of alcalase to the epoxy-activated resin, the remaining active groups were blocked with 3m glycine and the enzyme preparation was washed with an excess amount of phosphate-buffered saline. The second immobilization method used an amino-activated beaded resin (toyopearl af-amino-650m, tosoh bioscience llc, king of prussia, pensylvania, u.s.a.) and the linker 1,4-phenylene diisothiocyanate (pditc, sigma-aldrich, st. Louis, missouri, u.s.a.) as described elsewhere27. The tubes used for sample collection-reaction were made of glass. For the immobilization of alcalase to the inner surface of the glass tube, a linking process for successful functionalization of the silica-containing surface was employed using 3 aminopropyltriethoxysilane (santa cruz biotechnology, inc., dallas, texas, u.s.a.) prior to coupling the enzyme to the surface as described elsewhere27.
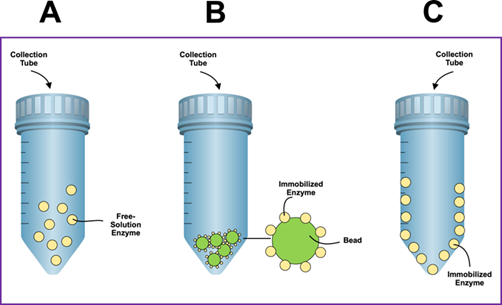
Figure 3. Diagrammatic representation of collection-reaction tubes where the sputum was disrupted, extracted, and digested to release its content using the nonionic detergent triton x-100 and the endonuclease alcalase. The tubes feature screw caps that provide a tight and secure seal. Panel a depicts the action of the protease on sputum as a free- solution enzyme in conjunction with triton x-100. Panel b depicts the action of the protease on sputum as an immobilized enzyme to beads used as solid support. Panel c depicts the action of the protease on sputum as an immobilized enzyme to the inner surface of a collection-reaction tube used as a solid support. The tube can be made of glass or polymeric material with a modified surface to attach one or more digestive enzymes.
Impressively, since the first days when some of the symptoms of covid-19 were shown in an apparent infected person, the presence of the sars-cov-2 virus was reported negative using the lfia testing for all specimens assayed, including nasopharyngeal, oropharyngeal, buccal, saline mouth rinse-gargle, and saliva, except when testing a sputum specimen in which the results were reported positive. Is the viral load in sputum detectable for 3 days, 5 days, or more, when almost all symptoms have disappeared? To answer this question, it was necessary to employ the modified disruption-extraction sample preparation protocol and search for the presence of the sars-cov-2 virus in sputum samples several days beyond initial testing. When the symptoms no longer persisted. As shown in (figure 4), a positive result was observed for the presence of sars-cov-2 virus tested in sputum specimens several weeks after obtaining expectorated sputum. As the frequency of coughing decreased with time, the amount of spontaneous sputum diminished as well. Nonetheless, the quantity of sputum obtained for 15 weeks was sufficient to perform the experiments. To make sure all experimental conditions were consistent for the 15-week experiments, strict protocols were maintained including the use of the same lfia covid-19 antigen-based test kits (flowflex covid-19 antigen home test, acon laboratories, inc., san diego, california, u.s.a.). Since the intensity of the colored band diminished in time, all tested samples were incubated for 2 hours to ensure the visibility of the band. Regarding the patient, there were no clinical manifestations presented after the initial 3 days of mild symptoms. Everything seemed to be normal during the rest of the experiments, except for an occasional coughing that persisted beyond the time the experiments were performed.
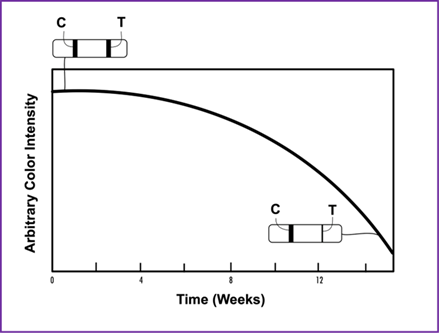
Figure 4. Illustration of a time experiment for the presence of sars-cov-2 virus in sputum samples obtained during a period of 15 weeks using the modified disruption-extraction sample preparation protocol. The color intensity of the test (t) line diminishes as the persistence of the coughing in the patient lessens and the amount of sputum collected is reduced as well. The sputum specimen was obtained from a patient that had covid-19-like symptoms for approximately 4-5 days but was asymptomatic for the rest of the 15-day period of testing, except for some minor coughing.
Preliminary experiments using alcalase immobilized to the surface of a beaded resin were performed (figure 5). The size of the beads used for immobilizing the protease was 65 microns, limiting the surface area for linking the proteolytic enzyme. Nonetheless, the procedure worked demonstrating the presence of viral entities in the sputum. Increasing the surface area for enzyme immobilization using smaller bead size may enhance color band intensity.
Figure 5. Diagrammatic representation of the sandwich format of lateral flow immunoassay test using the modified disruption-extraction sample preparation protocol for sputum. The disrupted-lysed sputum specimen was incubated for 2 hours at 25-degrees celsius in the triton x-100 solution containing beads with immobilized alcalase.
Discussion
Timely and reliable testing is important in controlling the covid-19 pandemic. Current covid-19 antigen testing is primarily carried out by obtaining a specimen via nasopharyngeal/oropharyngeal (np/op) swab samples and performing a rapid lateral flow immunoassay (lfia) disposable micro-device or related immunoassays. Nonetheless, these np/op screening tools do not rule out a covid-19 infection28. Furthermore, although np sampling is considered safe, single case reports and clinical observations indicate the possibility of several complications29.
Numerous studies have examined the presence of covid-19 in samples tested beyond the traditional positivity window expected in np/op rapid antigen-based and rt-pcr tests. For example, it is known that in sars-cov-2, viral rna is still present in feces of more than 60% of patients, after nasopharyngeal swab testing turned negative results by rt-pcr assays, suggesting that fecal-oral transmission may serve as an alternative route for sars-cov-2 transmission30-32. Another study found that at day 111 from a covid-19 patient’s initial testing, cytopathic effects were observed in specimens from a nasopharyngeal swab and sputum inoculated into veroe6/tmprss2 cells, and viral rna was detected in the culture supernatant by quantitative rt-pcr33. The centers for disease control and prevention recommend that patients infected within the past 90 days without new covid-19 symptoms should not be retested34.
Due to the complexity and viscosity of sputum, and the labor intense pretreatment of mucus specimens, various attempts have been made by several investigators to disrupt sputum samples16, including treatment of the collected sputum with mucolytic agents or reducing agents such as dithiothreitol (dtt), dithioerythritol (dte), tris(2-carboxyethyl)phosphine (tcep), or n-acetyl-l-cysteine (nalc) to reduce disulfide bonds of the oligomeric gel-forming mucins35,36. Other investigators have added hydrogen peroxide (h2o2) to react with endogenous catalase within the sputum to geneate oxygen. The o2 bubbles formed during the enzymatic reaction liquefy the sample without any additional instrumentation37. Detergents have also been used to disrupt all kinds of cellular and subcellular membranes38, and in Conjunction with solvents to inactivate viruses39. Furthermore, mechanical dissociation has been used to liberate cells from sputum40.
Apparently, increased use of sputum samples for the detection of the sars-cov-2 virus may be one way to answer this question, as well as many other queries that remain unanswered regarding the covid pandemic, including a greater understanding of the advantages and limitations of differential treatments, the degree of diagnostic accuracy and analytical sensitivity of certain rapid tests, in addition to some vaccine side effects41-45. A positive association between sputum viral load and disease severity for covid-19, as well as an increased risk of progression, has been reported46. In this study, the authors used sputum specimens instead of nasopharyngeal and oropharyngeal swabs because samples from the lower respiratory tract generally contain a higher level of viral load than nasopharyngeal and oropharyngeal swabs. Additionally, it has been reported that when testing sars-cov-2 rna in 1060 sputum samples, the rate of sample positivity in sputum was highest when compared with nasopharyngeal and oropharyngeal swab samples47. Furthermore, it has been demonstrated that patients with sars-cov-2 infection and severe covid-19 often have multiple coinfections, and their treatment is challenging48. Though sputum samples may provide further insight into the period of infectivity, severity of illness, and risk of progression of covid-19, sample processing methods must nevertheless improve to meet the yield of current nasopharyngeal rapid and rt-pcr testing.
Since the fundamental purpose of any diagnostic test is to help determine whether a patient has or does not have a particular condition, it was essential to corroborate the validity of the negative results reported by an urgent care clinic for the presence of sars-cov-2 antigen and rna viral constituents. Based on the clinical examination on the patient and on the negative laboratory result tests using np/op collected swab samples, the attending physician at the clinic concluded that the symptoms reported by the patient may have been related to a seasonal flu rather than to covid-19. On the other hand, as reported in this paper, using the patient’s sputum sample in conjunction with the modified pretreatment procedure for releasing the sputum constituents, it was demonstrated that the rapid lfia test yielded positive results for the presence of antigenic sars-cov-2 proteins. Such positive test results, suggest that the patient may have had covid-19 at the time of being tested.
The main limitation of our pilot study is the sample size, which includes only a single patient. Notably, repeated testing of antigenic protein was conducted in this patient for a total of 15 weeks that yielded consistent results. It is important to note that additional analysis of sars-cov-2 rna by rt-pcr yielded invalid results in two collected sputum samples that were kept in the freezer in the presence of triton-x before testing. As it has been known that pcr inhibitors are a very heterogeneous group of chemical substances49, it is possible that the high concentration of triton x-100 (1%) added to the sputum sample, or other constituents released from sputum may have contributed to the invalidity of the test. Another limitation of our pilot study is the lack of a respiratory viral pathogen panel that may have truly ruled out the presence of another respiratory infection. It is possible that the repeated positive test results could have been explained by the presence of another viral protein in the hydrolyzed sputum which may have caused cross reactivity, if there was any. In fact, it has been shown that sars-cov-2 shares homology and cross- reacts with vaccines, other viruses, common bacteria, and many human tissues50. Despite these limitations, the results of this study warrant further studies on larger scale to make this important observation of positive sputum-based lfia testing in an otherwise negative nasopharyngeal-based lfia test clinically significant.
Conclusion
A simple, rapid, non-invasive, and cost-effective pilot study method to study the sars-cov-2 antigenic proteins in sputum is described. Using commercially available lateral flow immunoassay kits, and inexpensive reagents such as detergents and proteolytic enzymes, it was possible to challenge a negative nasopharyngeal rapid antigen-based testing for covid-19 with a positive testing. This method may simplify the current tedious protocols which make use of silica-based columns, as well as strong and hazardous chemicals to extract biomolecules and to breakdown complex structures, such as dithiothreitol, guanidinium isothiocyanate, phenol-chloroform, and others. Pre-packet collection-reaction tubes with immobilized alcalase enzyme and detergent would be a simpler, safer, and distinctly efficient method to disrupt the complex sputum meshwork and yield a more accurate test for sars-cov-2 viral entities and other microbes or molecules of interest. Although several papers have reported the presence of sars-cov- 2 rna in sputum5-8,51, and a few have reported the use of target proteomics52 and surface-enhanced raman spectroscopy53 for the detection of sars-cov-2 antigens, to the best of our knowledge this is the first time that the presence of sars-cov-2 antigenic proteins has been reported in a disrupted/lysed sputum sample, using a detergent in combination with a proteolytic enzyme and a simple lateral flow immunoassay technology used as a detection assay platform.
Author contributions
N.a.g. conceived, designed, performed the experiments, and wrote the paper; d.e.g. analyzed the data and critically revised the manuscript. The authors have read and agreed with the final version of the manuscript.
Orcid numbers
N.a.g.: 0000-001-5504-376x d.e.g.: 0000-0002-5771-0586
Competing interest
N.a.g. is the inventor of patents pending on this subject.
Funding
This work did not receive any specific grant from funding agencies in the public, commercial, or not-for- profit sectors.
Compliance with ethical standards. Biological samples were obtained from one volunteer, from the start of the symptoms until the last experiment that lasted 15 weeks, with written and informed consent. The dated and signed consent document has been archived. A minimal risk study was approved by the ethical committee at princeton biochemicals, inc., and conducted according to ethical principles of the declaration of helsinki and good practice guidelines. The ethical approval was obtained from the institutional review board (approval number: pbi-03282023).
Safety statement
No unexpected or significant safety hazards are associated with the reported work.
Data availability statement
All data generated or analyzed during the study are included in the article.
References
2. ali m, han s, gunst cj, lim s, luinstra k, smiela m. Throat and nasal swabs for molecular detection of respiratory viruses in acute pharyngitis. Virol j 2015;12:178.
3. saleem sm, bhattacharya s. Sputum testing as the new mass screening method for covid-19 patients in india - a public health perspective. Int j prev med 2022;13:86.
4. pircalabioru gg, iliescu fs, mihaescu g, et al. Advances in the rapid diagnostic of viral respiratory tract infections. Front cell infect microbiol 2022;12:807253.
5. han h, luo q, mo f, long l, zheng w. Sars-cov-2 rna more readily detected in induced sputum than in throat swabs of convalescent covid-19 patients. Lancet infect dis 2020;20(6):655-656.
6. qu ym, kang em, cong hy. Positive results of sars-cov-2 in sputum from a cured patient with covid-19. Travel med infect dis 2020 34:101619
7. jeong jh, kim kh, jeong sh, park jw, lee sm, seo yh. Comparison of sputum and nasopharyngeal swabs for detection of respiratory viruses. J med virol 2014;86(12):2122-2127.
8. branche ar, walsh ee, formica ma, falsey ar. Detection of respiratory viruses in sputum from adults by use of automated multiplex pcr. J clin microbiol 2014;52(10):3590-3596.
9. leal j, smyth hdc, ghosh d. Physicochemical properties of mucus and their impact on transmucosal drug delivery. Intern j pharm 2017;532(1):555-572.
10. kratochvil mj, kaber g, demirdjian s, et al. Biochemical, biophysical, and immunological characterization of respiratory secretions in severe sars- cov-2 infections. Jci insight 2022;7(12):e152629.
11. d’amato m, iadarola p, viglio s. Proteomic analysis of human sputum for the diagnosis of lung disorders: where are we today? Int j mol sci 2022;23(10):5692.
12. fahy jv, dickey bf. Airways mucus function and dysfunction. N engl j med 2010;363:2233-2247.
13. lai t, xiang f, zeng j, et al. Reliability of induced sputum test is greater than that of throat swab test for detecting sars-cov-2 in patients with covid-19: a multi-center cross-sectional study. Virulence 2020;11(1):1394-1401.
14. cukic v. The most common detected bacteria in sputum of patients with acute exacerbation of copd. Mater sociomed 2013;25(4):226-229.
15. blakeborough l, watson js. The importance of obtaining a sputum sample and how it can aid diagnosis and treatment. Br j nurs 2019;28(5):295-298.
16. park hj, woo a, cha jm, lee ks, lee my. Closed-type pre-treatment device for point-of-care testing of sputum. Sci rep. 2018;8:16508.
17. guzman na, guzman de. Advancements in portable instruments based on affinity-capture-migration and affinity-capture-separation for use in clinical testing and life science applications. J chromatogr a 2023;1704:464109.
18. hsieh wy, lin ch, lin tc, et al. Development and efficacy of lateral flow point-of-care testing devices for rapid and mass covid-19 diagnosis by the detections of sars-cov-2 antigen and anti-sars-cov-2 antibodies. Diagnostics 2021;11(10):1760.
19. zhang y, chai y, hu z, et al. Recent progress on rapid lateral flow assay-based early diagnosis of covid-19. Front bioeng biotechnol 2022;10:866368.
20. koczula km, gallotta a. Lateral flow assays. Essays biochem 2016;60(1):111-120.
21. baptista p, pereira e, eaton p, et al. Gold nanoparticles for the development of clinical diagnosis methods. Anal bioanal chem 2008;391(3):943- 950.
22. shen f, sergi c. Sputum analysis. Statpearls 2022.
23. datta s, christena lr, rajaram yrs. Enzyme immobilization: an overview on techniques and support materials. 3 biotech 2013;3(1):1-9.
24. wong ls, khan f, micklefield j. Selective covalent protein immobilization: strategies and applications. Chem rev 2009;109(9):4025-4053.
25. hodneland cd, lee ys, min dh, mrksick m. Selective immobilization of proteins to self- assembled monolayers presenting active site-directed capture ligands. Proc natl acad sci usa 2002;99(8):5048-5052.
26. mateo c, torres r, fernández-lorente g, et al. Epoxy-amino groups: a new tool for improved immobilization of proteins by the epoxy method. Biomacromolecules 2003;4(3):772-777.
27. guzman na, trebilcock ma, advis jp. The use of a concentration step to collect urinary components separated by capillary electrophoresis and further characterization of collected analytes by mass spectrometry. J liq chromatogr 1991;4(5):997-1015.
28. winichakoon p, chaiwarith r, liwsrisakun c, et al. Negative nasopharyngeal and oropharyngeal swabs do not rule out covid-19. J clin microbiol 2020;58(5):00297-20.
29. koskinen a, tolvi m, jauhiainen m, kakäläinen e, laulajainen-hongisto a, lamminmäki s. Complications of covid-19 nasopharyngeal swab test. Jama otolaryngol. Head neck surg 2021;147(7):672-674.
30. chen y, chen l, deng q, et al. The presence of sars-cov-2 rna in the feces of covid-19 patients. J med virol 2020;92(7):833-840.
31. chen c, gao g, xu y, et al. Sars-cov-2-positive sputum and feces after conversion of pharyngeal samples in patients with covid-19. Ann inter med 2020;172(12)832-834.
32. jones dl, baluja mq, graham dw, et al. Shedding of sars-cov-2 in feces and urine and its potential role in person-to-person transmission and the environment-based spread of covid-19. Sci total environ 2020;749:141364.
33. abe t, ikeda t, tokuda y, et al. A patient infected with sars-cov-2 over 100 days. Qjm an int j med 2021;114(1):47-49.
34. coffey kc, diekema dj, morgan dj. Interpreting sars-cov-2 test results. J am med assoc 2021;326(15):1528-1529.
35. nicholas b, skipp p, mould r, et al. Shotgun proteomic analysis of human-induced sputum. Proteomics 2006;6:4390-4401.
36. kang a, yeom m, kim h, et al. Sputum processing method for lateral flow immunochromatographic assays to detect coronaviruses. Immune netw 2021;21(1):e11.
37. clemente a, alba-patiño a, santopolo g, et al. Immunodetection of lung igg and igm antibodies against sars-cov-2 via enzymatic liquefaction of respiratory samples from covid-19 patients. Anal chem 2021;93(12):5259-5266.
38. guzman na, cutroneo kr. Association of prolyl hydroxylase activity with membranes. Biochem biophys res commun 1973;52(4):1263-1270.
39. rabenau hf, biesert l, schmidt t, bauer g, cinati j, doerr hw. Sars-coronavirus (sars-cov) and the safety of a solvent/detergent (s/d) treated immunoglobulin preparation. Biologicals 2005;33(2):95-99.
40. barber c, lau l, ward ja, et al. Sputum processing by mechanical dissociation: a rapid alternative to traditional sputum assessment approaches. Clin respir j 2021;15(7):800-807.
41. zhang c, huang s, zheng f, dai y. Controversial treatments: an update understanding of the coronavirus disease 2019. J med virol 2020;92(9):1441-1448.
42. qorib m, oladunni t, denis m, ososanya e, cotae p. Covid-19 vaccine hesitancy: text mining, sentiment analysis and machine learning on covid-19 vaccination twitter dataset. Expert syst appl 2023;212:118715.
43. juniastuti, furqoni ah, amin m, et al. The evaluation results of proposed antigen rapid diagnostic tests for covid-19: some possible factors might influence. Infection 2023;2:1-7.
44. piek a, knoester m, gard l, lokate m, niesters hgm, van leer-buter cl. Sars-cov-2 antigen tests for screening of healthcare workers; experience with over 48,000 combined antigen tests and rt-pcr tests. J clin virol 2023;158:105326.
45. wang m, liao z. Sars-cov-2 and covid-19: how much do we know? Acta virol 2020;64(3):288-296.
46. yu x, sun s, shi y, wang h, zhao r, sheng j. Sars-cov-2 viral load in sputum correlates with risk of covid-19 progression. Crit care 2020;24:170.
47. mohammadi a, esmaeilzadeh e, li y, bosch rj, li jz. Sars-cov-2 detection in different respiratory sites: a systematic review and meta-analysis. Ebiomedicine 2020;59:102903.
48. fujimoto d, fukuya m, terao s. Sputum characteristics of patients with severe covid-19: report of two cases with immunocytochemical detection of sars-cov-2 spike protein. Med mol morphol 2022;55(4):316-322.
49. schrader c, schielke a, ellerbroek l, johne r. Pcr inhibitors - occurrence, properties and removal.
J
appl microbiol 2012;113(5):1014- 1026.
51. chen c, gao g, xu y, et al. Sars-cov-2-positive sputum and feces after conversion of pharyngeal
samples in patients with covid-19. Ann int med 2020;172(12):832- 834.
52. bezstarosti k, lamers mm, doff was,
et al. Targeted proteomics as a tool to detect
sars-cov-2 proteins in clinical specimens. Plos one 2021;16(11):e0259165.
53. huang j, wen j, zhou m, et al. On-site detection of sars-cov-2
antigen by deep learning-based surface-enhanced raman spectroscopy and its
biochemical foundations. Anal chem 2021;93:9174-9182.