Leveraging Artificial Intelligence (AI) in Finding the Interconnected Epidemics and Genetic Predisposition of Obesity and Type 2 Diabetes: A Review
Abstract
Keywords: Obesity; Type2 Diabetes mellitus; Risk factors; Artificial intelligence, Genetic predisposition.
Introduction
Obesity
and type 2 diabetes mellitus (t2dm) represent two of the most pressing health
challenges globally, often referred to as twin epidemics due to their strong
interrelationship. The rising prevalence of these conditions has significant
implications for public health, healthcare systems and socio-economic
stability. This editorial examines the intricate links between obesity and
t2dm, exploring the pathophysiological mechanisms, risk factors and potential
intervention strategies. The intersection of genetics with obesity and t2dm
underscores the complex etiology of these metabolic disorders. Genetic
predisposition plays a crucial role in influencing an individual's risk of
developing both obesity and t2dm. This predisposition is mediated through
various gene variants that regulate body weight, energy homeostasis and insulin
sensitivity.
The
link between obesity and type 2 diabetes
The
elaborate connections and sharing of pathophysiological mechanisms between
obesity and t2dm are well-established. Both type 2 diabetes and obesity include
excess body fat, particularly visceral fat, contributing to insulin resistance
and nafld (non-alcoholic fatty liver disease) and a constellation of metabolic
abnormalities in obese individuals a hallmark of t2dm. However, in some
individuals, t2dm can also occur inversely before obesity with inherent insulin
resistance resulting in increased hepatic glucose production and elevated
insulin levels, which are the actual cause of obesity1,2. Genetic and environmental factors play
an important role in connecting obesity and t2dm and these factors involved in
low insulin secretions from b-cells and peripheral insulin resistance, leading
to elevated levels of fatty acids. This causes a decrease in glucose transport
into muscle cells, increased fat breakdown and hepatic glucose production. This
ectopic fat breakdown interferes with insulin signaling pathways, resulting in
insulin resistance, where cells fail to respond effectively to insulin3. Further, adipose tissue, particularly in
obese individuals, secretes pro-inflammatory cytokines like interleukin-6
(il-6) and tumor necrosis factor-alpha (tnf-α), interleukin-1β (il-1β),
monocyte chemoattractant protein-1 (mcp-1), il-6 and others released by
activated tissue macrophages and by adipocytes play a significant role. These
cytokines provoke important responses in the liver, skeletal muscle, fat tissue
and pancreas, resulting in endocrine dysfunction, impaired glucose disposal,
impaired β-cell function and reduced suppression of glucose production. In
addition to cytokines, adipose tissue produces various adipokines such as
leptin and adiponectin. In obesity, an imbalance in these adipokines occurs,
with increased leptin (leptin resistance) and decreased adiponectin levels,
contributing to the pathogenesis of t2dm. The flow chart (figure 1) by depicts the role
of insulin sensitivity leading to obesity and t2dm4.
Obesity induces er stress in various tissues, leading to the activation of
stress signalling pathways that impair insulin action. This contributes to the
development and progression of t2dm (figure 1)
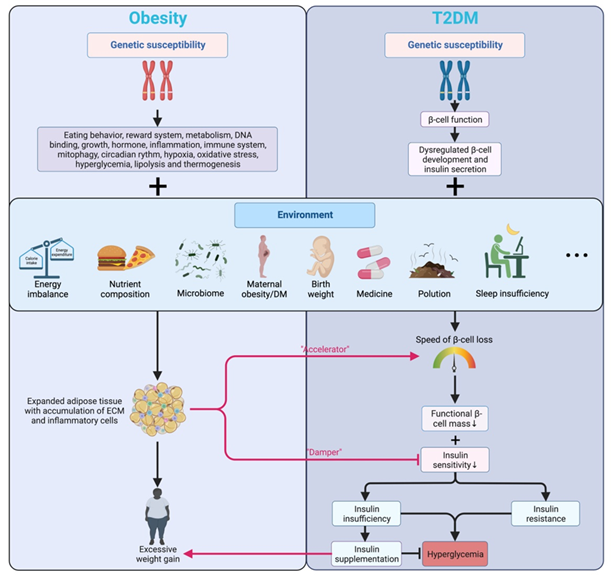
Risk factors for obesity and t2dm
Table 1: the risk factors for obese individuals who may develop type 2 diabetes mellitus
Role
of ai in t2dm and obesity
In the past years, several studies have
been developed involving information technology (it) in the healthcare industry
to manage patient data and to improve diagnosis and treatment with artificial
intelligence (ai). This specific area of ai of computer science is also a field
in science and engineering and it has provided reasoning tools to support
clinical decision-making for healthcare professionals16,17.
Moreover, incorporating ai-based technologies into medical practice is expected
to produce substantial changes in many areas of medicine and healthcare18,19. Ai
can help identify risk factors for diabetes development, addressing human
limitations and biases when working with massive datasets. Diabetes prevention
strategies can be tailored to specific individuals by identifying modifiable
risk factors. By incorporating ai analysis a large dataset of demographic
details and clinical parameters such as age, gender, bmi, and other
socioeconomic status helps in predicting the risk factors of obesity and t2dm.
On the other hand, clinical parameter interpretation test results help find
accurate and customized treatment plans for patients using ai technology. The
development of newer technologies in managing the healthcare system is a
potentially significant reduction in the burden of obesity and t2dm treatment
which helps in a more proactive and personalized approach20.
Artificial intelligence and genomics
After the completion of human genome
projects, the genomes from multiple human populations and diverse primate and
non-primate animals have been sequenced and stored in public databases,
allowing for key discoveries through dna sequence comparisons and evaluations
of allele frequencies and identifying nucleotide variation in different groups
are listed in (table 2)21,22.
Table 2: artificial intelligence-
power tool for genome sequencing aspects
|
Aspects |
Description |
|
accelerated sequencing |
Ai shortens the time and
expense associated with genome sequencing. |
|
Error reduction |
Ai decreases errors, increasing
the accuracy of genome sequencing. |
|
variant identification |
Ai swiftly and correctly
pinpoints genetic variations related to diseases or traits. |
|
Personalized medicine |
Ai uses genomic data analysis
to personalize medicines based on each patient's genetics. |
|
Population studies |
Large-scale datasets are
analyzed using ai to provide insights about population-level genetic
variants. |
|
Structural variation analysis |
Large-scale genomic
rearrangements and structural changes can be found with the help of ai. |
|
Data integration |
Ai combines clinical,
environmental, lifestyle, and genomic data to provide thorough insights. |
|
Ethical considerations |
Sensitive
genomic data storage and dissemination provide ethical difficulties. Aspects description accelerated sequencing ai shortens the time
and expense associated with genome sequencing. Error
reduction ai decreases errors, increasing the accuracy of genome sequencing. variant identification ai swiftly and
correctly pinpoints genetic variations related to diseases or traits. Personalized
medicine ai uses genomic data analysis to personalize medicines based on each
patient's genetics. Population
studies large-scale datasets are analyzed using ai to provide insights about
population-level genetic variants. Structural
variation analysis large-scale genomic rearrangements and structural changes can be
found with the help of ai. Data integration ai combines
clinical, environmental, lifestyle, and genomic data to provide thorough
insights. Ethical
considerations sensitive genomic data storage and dissemination provide ethical
difficulties. |
Over the last two decades,
significant progress has been achieved in defining protein sequence variation
and function (proteomics), as well as rna transcript isoforms and their
expression patterns23. Parallel,
artificial intelligence is an essential tool in genetic analysis. Genomic
analysis employs artificial intelligence (ai) methods such as machine learning
(ml) and deep learning (dl) to analyze and understand enormous volumes of
genetic data (figure 2). These
algorithms may use enormous datasets to identify patterns, predict the
outcomes, and categorize genetic variations based on training from large
datasets.
Figure 2: big genomic data analysis requires artificial intelligence (ai) methods such as machine learning (ml) and deep learning (dl).
Artificial intelligence and their
roles in snps analysis
As mentioned earlier, the
application of artificial intelligence tools to determine which single
nucleotide polymorphisms (snps) influence disease development is one of the
features of medical research, as the use of ai techniques may potentially help
physicians to identify the early diagnosis of snps related to type 2 diabetes
and obesity and to build a decision support tool for risk prediction.
Therefore, the data generated by ai for genetic predisposition plays a
significant role in determining an individual's susceptibility to both obesity
and t2dm. Certain gene variants influence body weight regulation and insulin
sensitivity. To date, genetic techniques have provided evidence that some
genetic variants have a role in obesity and insulin sensitivity in t2dm. Some
of them were cohort studies and investigated a large number of loci and snps, some
of them are listed in (table 3) and
described below. The results of the studies showed a significant association of
genetic variants with the major markers of t2d and obesity24.
Some of genes directly related to obesity and t2dm are described below:
Table
3:
important key genes involved in obesity and type 2 diabetes mellitus (t2dm)
|
Gene |
Associated disease |
Function/role |
|
Fto |
Obesity,
t2dm |
fat mass
and obesity-associated gene; influences body mass and fat accumulation. |
|
Tcf7l2 |
T2dm |
transcription
factor 7-like 2; involved in insulin secretion and glucose production. |
|
Lep |
Obesity |
leptin
gene; regulates appetite and energy balance. |
|
Lepr |
Obesity |
leptin
receptor gene; mediates the effects of leptin |
|
Pparg |
Obesity,
t2dm |
|
|
Kcnj11 |
T2dm |
potassium
inwardly-rectifying channel, subfamily j, member 11; involved in insulin
release. |
|
Abcc8 |
T2dm |
atp-binding
cassette transporter sub-family c member 8; regulates insulin secretion. |
|
Mc4r |
Obesity |
melanocortin
4 receptor; involved in energy homeostasis and appetite regulation. |
|
Irs1 |
T2dm |
insulin
receptor substrate 1; plays a role in insulin signalling. |
|
Adipoq |
Obesity,
t2dm |
Adiponectin
gene; involved in glucose regulation and fatty acid breakdown. |
Fto gene: the fat mass and obesity-associated (fto) gene is one of the most studied genes linked to obesity and t2dm. Variants in the fto gene, particularly single nucleotide polymorphisms (snps) like rs9939609, are associated with increased body mass index (bmi) and higher fat mass. Individuals with certain fto variants exhibit a stronger appetite, reduced satiety and a preference for high-calorie foods, contributing to weight gain and obesity. This gene is also implicated in the regulation of adipocyte function, influencing insulin sensitivity and the risk of t2dm.in a study conducted25, it was observed that individuals carrying the risk allele (a allele) of the rs9939609 snp in the fto gene had 1.67 times higher odds of developing obesity compared to those without the risk allele. Furthermore, another study demonstrated that this variant was associated with increased t2dm risk, independent of bmi, highlighting its direct impact on metabolic pathways26.
Tcf7l2 gene: the transcription factor 7-like 2 (tcf7l2) gene is among the strongest genetic determinants of t2dm. Variants like rs7903146 have been consistently associated with increased t2dm risk, independent of obesity. Tcf7l2 influences insulin secretion by affecting the function of pancreatic beta cells. Individuals with high-risk variants of tcf7l2 often exhibit impaired insulin secretion, leading to hyperglycemia and the subsequent development of t2dm.a genome-wide association study (gwas) identified the tcf7l2 variant rs7903146 as a significant predictor of t2dm27. Carriers of this variant had up to a 1.5-fold increased risk of t2dm, with the effect being more pronounced in individuals with a family history of the disease, highlighting the gene's critical role in genetic predisposition.
Lep: leptin plays an important role in regulating adipose-tissue mass in obese individuals. It has been shown that leptin is over-expressed at the gene level in the adipose tissue of individuals with obesity28. Moreover, other studies showed that strong positive associations exist between plasma leptin levels and body fat percentage29,30. The single nucleotide polymorphism (snp) in the leptin gene, especially rs7799039 g > a snp, was implicated in the development of obesity31. A recent report on snp suggested the association of leptin promoter gene variations i.e., rs72563764c>t and rs7799039g>a with both diabetes and obesity32. A few research reports are available on the association between human leptin gene variants and obesity traits in india33,34. In vivo, experimental studies confirmed a polymorphism in the ob gene. This confirmation of polymorphism alters the leptin protein function such that mice become morbidly obese35.
Lepr gene: leptin has its physiological action via attaching to the leptin receptor, which is a single transmembrane protein in the class i cytokine receptor family. Upon leptin binding to its leptin receptor, it activates various it activates multiple intracellular signaling pathways, including: including the jak2/stat5 pathway, the extracellular signal-regulated kinase (erk) pathway, and the insulin receptor substrate (irs)-phosphoinositide 3-kinase (pi3k)/akt pathway, which diverges to separate downstream signaling 36,37. The gene lepr encoding leptin receptor on the chromosomes 1, respectively. The lepr has been linked to the presence of single-nucleotide polymorphisms (snps) that may modulate the circulating concentration of this adipokine38. The lepr genes have been associated with obesity, deregulation of blood pressure levels, and sympathetic hyperactivity39,40. The most studied snps, rs1137101 (+668a > g) are situated in the exon 6 region of lepr. A study from illangasekera et al, demonstrates that the presence of the variant `g allele’ of the lepr q223r polymorphism is associated with greater bmi and wc measures41. Similar results were also reported by becer et al., where the association between the `g’ allele and obesity related anthropometric measures was observed only in obese subjects42. Furthermore, a cohort research study identified that an elevated level of lepr expression is linked to increased neoangiogenesis and metastatic potential in colorectal cancer survival (crc), whereas low lepr expression correlates with modest rates of proliferation43.
Pparg gene: the peroxisome proliferator-activated receptor gamma (pparg) gene is critical in adipocyte differentiation and lipid metabolism. The pro12ala variant (rs1801282) of the pparg gene is associated with a reduced risk of t2dm, possibly due to its effects on improving insulin sensitivity44. This variant is believed to alter the gene’s activity, leading to more favorable fat storage and utilization patterns that protect against insulin resistance and t2dm. The diabetes prevention program (dpp) study examined the role of the pparg pro12ala variant in response to lifestyle interventions and metformin45. The study found that individuals with the ala allele had a lower incidence of t2dm, particularly when combined with lifestyle interventions, suggesting a gene-lifestyle interaction in preventing diabetes onset.
Kcnj11: the synthesis of insulin and secretion is regulated by a family of potassium channels including potassium inwardly rectifying channels, subfamily, member 11(kcnj11). The kcnj11 gene, located 4.5 kb from abcc8 on chromosome 11p15.1, contains a single exon encoding the 390 amino acid kir6.2 protein. In kcnj11 gene several single nucleotide polymorphisms several single nucleotide polymorphisms have been identified, which may alter mrna and protein expression (rs5219, rs2237892, and rs151290)46. A recent study has shown that there is an association of genetic variants of kcnj11 genes with the risk of type 2 diabetes mellitus (t2dm) in the indian population47. Another study suggested that single nucleotide polymorphism of kcnj11 (rs5219) gene is associated with glycemic status and insulin resistance and pregnant women with t allele were at higher risk of developing gestational diabetes mellitus.
Abcc8: the abcc8 gene, which encodes the sulfonylurea receptor (sur), is the regulatory subunit of the katp channel and plays an important role in insulin production48. Research in a french adult type 2 diabetes outpatient cohort of 139 individuals discovered two (1.5%) potentially causal abcc8 mutations49. Another investigation in a large cohort of nonobese individuals with a diagnostic age < 40 years and a family history of diabetes showed 8 (8/1564, 0.5%) abcc8 mutations50. In addition, an east asian investigation discovered one abcc8 variation (0.9%) among 109 putative monogenic diabetes patients (park et al., 2019. Various studies estimate the frequency of abcc8 mutations to be between 0.5% and 1.5%. It implies that the beta-cell protein sulfonylurea receptor (sur1's) subunit atp-sensitive potassium (katp) channel, which encodes the abcc8 gene, is responsible for a small portion of neonatal diabetes mellitus (nndm).
Mc4r gene: the melanocortin 4 receptor (mc4r) gene is another critical gene influencing body weight regulation. Mutations in mc4r are the most common genetic cause of monogenic obesity, affecting appetite control and energy expenditure. Individuals with mc4r mutations tend to have severe early-onset obesity and are at a higher risk for t2dm due to the associated insulin resistance51. Found that mc4r mutations were present in approximately 6% of individuals with severe obesity. These individuals also had a higher incidence of t2dm, illustrating the direct impact of mc4r on both obesity and metabolic dysregulation.
Irs1: insulin receptor substrate (irs) molecules are important mediators of insulin signaling. The irs-1 is the first member of the family to be identified. The irs-1 gene, located on chromosome 2, has both the 5′-untranslated region and the protein-coding region in a single exon. Several single nucleotide polymorphisms in the irs genes have been found, but only irs-1's gly to arg 972 alteration appears to have a pathogenic role in the development of type 2 diabetes mellitus52,53. Previous studies have shown that the irs-1 gene's frequent polymorphism (rs 1801278) is a glycine to arginine substitution (ggg ↔ agg) at codon 972 (g972r), which may contribute to type 2 diabetes through insulin resistance and decreased secretion54,55. A recent study by bedair et al., 2021 suggested that irs-1 g972r (rs 1801278) polymorphism might be a contributing risk factor for the development of type 2 dm56. The study suggested that mutant allele (a) of irs-1 polymorphism is a risk factor for type 2 diabetes mellitus even in subjects with normal body weight. The increase in body mass index may be an independent risk factor for the development of type 2 diabetes mellitus.
Adipoq: adiponectin is a protein expressed and secreted by adipose tissues. Adiponectin gene (adipoq) locus, 3q27, has been strongly linked to various metabolic disorders like- impaired glucose tolerance, t2d, obesity and dyslipidemia57. Genetic research discovered that additive genetic factors might explain 80% of the variation in blood adiponectin levels among nonobese individuals. Recent studies on different ethnic groups have shown the positive association of certain snps of the adipoq gene in t2dm58,59. Rs266729 is an adipoq gene snp that is thought to regulate promoter activity. In a study of 1004 adult obese adults, the g allele of rs266729 was linked to decreased blood adiponectin levels and an increased risk of hyperglycemia60. Whereas, as a cross-sectional study on mexican-mestizo individuals who carry the gg genotype have significantly higher levels of serum adiponectin than individuals who carry the tt or the tg genotypes of the snp rs2241766 adipoq gene61.
The key insights gained from the above data, emphasizes how these genes contribute to our understanding of the complex interplay between genetic predisposition and metabolic disorders. The implications of these findings pave the way for future research and point out the potential therapeutic targets, for personalized medicine. Moreover, the limitations of this study is that the gene list is not exhaustive and neither are the pathologies, however, the study suggests areas for further investigation to deepen our understanding of the genetic mechanisms underlying obesity and t2dm. Further, this article does not identify or propose the drugs for these two disorders as treatment options. Insulin a hormone, functions as a chemical messenger that regulates various physiological processes. It exerts its effects by binding to receptors on the cell surface, triggering a cascade of intracellular events through secondary messengers like camp. Thus, finding molecules (drugs) for its inhibitory effects to control diabetes is like altering gene expression which may or may not be as anticipated, especially in cases of comorbidities.
Conclusion
The relationship between obesity and t2dm is complex and
multifactorial, involving genetic, environmental, and behavioral factors. Given
the global burden of these conditions, an integrated approach that combines
individual-level interventions with public health strategies is essential and
discussed in this review. Chronic insulin resistance places a higher demand on
pancreatic β-cells to produce more insulin. Over time, this can lead to β-cell
exhaustion and dysfunction, reducing insulin secretion and contributing to
hyperglycemia. Future research should focus on understanding the molecular
mechanisms linking genetic basis of obesity to t2dm, which could lead to the
discovery of novel therapeutic targets. The genetic predisposition to obesity
and t2dm involves a complex interplay between multiple gene variants,
environmental factors and epigenetic modifications. Understanding the specific
genetic factors and the demographs of individuals may contribute to these
conditions and can form personalized prevention and treatment strategies,
particularly in individuals with a strong family history of obesity or t2dm.
Research should continue to explore using ai as a tool to unfold the epigenetic
mechanisms, as well as their interactions with lifestyle factors, to identify
more effective interventions for these interrelated diseases. Addressing these
twin epidemics requires a collaborative effort between ai, healthcare
providers, policymakers and communities to create environments that promote
healthy lifestyles and reduce the incidence of these interrelated conditions.
Declarations
Conflict of interest: the authors declare none.
Funding: no funds were received for this study.
Ethical issues: not required
Author contribution: all authors equally contributed and
agree to publish.
References
1. Mandrup poulsen t. Type 2
diabetes mellitus. A metabolic autoinflammatory disease dermatol clin
2013;31(3):495-506.
2. Eizirik dl, pasquali l,
cnop m. Pancreatic β-cells in type 1 and type 2 diabetes mellitus: different
pathways to failure nat rev endocrinol 2020;16(7):349-362.
3. Roder me, porte d, jr.
Schwartz rs, kahn se. Disproportionately elevated proinsulin levels reflect the
degree of impaired b cell secretory capacity in patients with
noninsulin-dependent diabetes mellitus. J clin endocrinol metab
1998;83(2):604-608.
4. Ruze r, liu t, zou x, song
j, chen y, xu r et al. Obesity and type 2 diabetes mellitus: connections in
epidemiology, pathogenesis and treatments. Front endocrinol (lausanne)
2023;21:1161521.
5. Kyrou i, randeva hs,
tsigos c, kaltsas g, and weickert mo. Clinical problems caused by obesity.
South dartmouth: endotext 2010.
6. Chen l, magliano dj,
zimmet pz. The worldwide epidemiology of type 2 diabetes mellitus[mdash]present
and future perspectives. Nat rev endocrinol 2012;8(4):228-236.
7. Gastaldelli a. Abdominal
fat: does it predict the development of type 2 diabetes? Am j clin nutr
2008;87(5):1118-1119.
8. Shutto y, shimada m,
kitajima m, yamabe h, razzaque ms. Lack of awareness among future medical
professionals about the risk of consuming hidden phosphate-containing processed
food and drinks. Plos one 2011;6(12):e29105.
9. Chao a, grilo cm, white
ma, sinha r. Food cravings, food intake, and weight status in a community-based
sample. Eat behav 2014;15(3):478-482.
10. Iyen b, vinogradova y,
akyea rk, weng s, qureshi n, kai j. Ethnic disparities in mortality among
overweight or obese adults with newly diagnosed type 2 diabetes: a
population-based cohort study. J endocrinol invest 2022;45(5):1011-1020.
11. Ali mk, narayan v, tandon
n. Diabetes & coronary heart disease: current perspectives. Indian j med
res 2010;132(5):584-597.
12. Wondmkun yt. Obesity,
insulin resistance, and type 2 diabetes: associations and therapeutic
implications. Diabetes metab syndr obes 2020;13:3611-3616.
13. Arneth, b. Mechanisms of
insulin resistance in patients with obesity. Endocrines 2024;5(2):153-165.
14. Ashraf h, laway ba, afroze
d, wani ai. Evaluation of proinflammatory cytokines in obese vs non-obese
patients with metabolic syndrome. Indian j endocrinol metab 2018;22(6):751-756.
15. Perry bd, caldow mk,
brennan-speranza tc, et al. Muscle atrophy in patients with type 2 diabetes
mellitus: roles of inflammatory pathways, physical activity and exercise. Exerc
immunol rev 2016;22:94-109.
16. Lisboa pj. A review of
evidence of health benefit from artificial neural networks in medical
intervention. Neural netw 2002;15(1):11-39.
17. Russell sj, norvig p.
Artificial intelligence: a modern approach. 3rd ed. Usa: pearson education inc
2010.
18. Roski j, chapman w, heffner
j, trivedi r, del fiol g, kukafka r et al. 'How artificial intelligence is
changing health and health care'. In artificial intelligence in health care:
the hope, the hype, the promise, the peril. Editors: matheny m, israni st,
ahmed m, whicher d. Washington, dc: national academy of medicine, 2019.
19. Fihn sd, saria s, mendonça
e, hain s, matheny m, shah n et al. 'Deploying ai in clinical settings. In
artificial intelligence in health care: the hope, the hype, the promise, the
peril', editors: matheny m, israni st, ahmed m, whicher d. Washington, dc:
national academy of medicine, 2019.
20. J. Huang, et al. Artificial
intelligence for predicting and diagnosing complications of diabetes j.
Diabetes sci. Technol 2023;17(1):224-238.
21. Margulies eh, & birney
e. Approaches to comparative sequence analysis: towards a functional view of
vertebrate genomes. Nature reviews genetics 2008;9(4):303-313.
22. Karczewski kj, francioli
lc, tiao g, cummings bb, alföldi j, wang q, et al. The mutational constraint
spectrum quantified from variation in 141,456 humans nature
2020;581(7809):434-443.
23. Jumper j, evans r, pritzel
a, green t, figurnov m, ronneberger o, er al. Highly accurate protein structure
prediction with alphafold nature 2021;596(7873):583-589.
24. Manolio ta. Genomewide
association studies and assessment of the risk of disease. N engl j med
2010;8,363(2):166-176.
25. Dina c, meyre d, gallina
s, et al. Variation in fto contributes to childhood obesity and severe adult
obesity. Nat genet 2007;39(6):724-726.
26. Frayling tm, timpson nj,
weedon mn, et al. A common variant in the fto gene is associated with body mass
index and predisposes to childhood and adult obesity. Science
2007;316(5826):889-894.
27. Grant sf, thorleifsson g,
reynisdottir i, et al. Variant of transcription factor 7-like 2 (tcf7l2) gene
confers risk of type 2 diabetes. Nat genet 2006;38(3):320-323.
28. Lonnqvist f, arner p,
nordfors l, schalling m. Overexpression of the obese (ob) gene in adipose
tissue of human obese subjects nat med 1995;1:950-953.
29. Hamilton bs, paglia d, kwan
ay, deitel m. Increased obese mrna expression in omental fat cells from
massively obese humans nat med 1995;1(9):953-956.
30. Considine rv, sinha mk,
heiman ml, kriauciunas a, stephens tw, nyce mr, et al. Serum
immunoreactive-leptin concentrations in normal-weight and obese humans. N engl
j med 1996;334(5):292
295.
31. Portolés o, sorlí jv,
francés f, coltell o, gonzález ji, sáiz c. Effect of genetic variation in the
leptin gene promoter and the leptin receptor gene on obesity risk in a
population-basedcase-control
Study in spain, european journal of epidemiology 2006;21(8):605-612.
32. Dar rubiya, rasool shabhat,
waza ajaz ahmad, ayoub gazalla, qureshi meenu, zargar abdul hamid et al.
Polymorphic analysis of leptin promoter in obese/diabetic subjects in kashmiri
population. Indian journal of endocrinology and metabolism 2019;23(1):111-116.
33. Dasgupta s, salman m,
siddalingaiah lb, lakshmi gl, xaviour d and sreenath j. Genetic variants in
leptin: determinants of obesity and leptin levels in south indian population,
adipocyte 2014;4(2):135-140.
34. Bains v, kaur h,
badaruddoza b. Association analysis of polymorphisms in lep (rs7799039 and
rs2167270) and lepr (rs1137101) gene towards the development of type 2 diabetes
in north indian punjabi population, gene 2020;754.
35. Igel m, becker w, herberg
l, joost hg. Hyperleptinemia, leptin resistance, and polymorphic leptin
receptor in the new zealand obese mouse endocrinology 1997;138:4234-4239.
36. Ahima rs, osei sy. Leptin
signaling. Physiol behav. 2004;81(2):223-241.
37. Kim sy, lim jh, choi sw,
kim m, kim st., kim ms et al. Preferential effects of leptin on cd4 t cells in
central and peripheral immune system are critically linked to the expression of
leptin
Receptor. Biochem. Biophysical res. Commun 2010;394(3):562-568.
38. Dam, j. Trafic et
signalisation du récepteur de la leptine [traffic and signalisation of the
leptin receptor]. Biol. Aujourdhui 2018;212:35-43.
39. Saad a, adam i, elzaki seg,
awooda ha, hamdan hz. Leptin receptor gene polymorphisms c.668a>g and
c.1968g>c in sudanese women with preeclampsia: a case-control study. Bmc
med.
Genet 2020;21:162.
40. Srinivasan g, parida s,
pavithra s, panigrahi m, sahoo m, singh tu et al. Leptin receptor stimulation
in late pregnant mouse uterine tissue inhibits spontaneous contractions by
increasing no and
Cgmp cytokine 2021;137:155341.
41. Illangasekera ya,
kumarasiri pvr, fernando dj, dalton df.
Association of the leptin receptor q223r (rs1137101) polymorphism with obesity
measures in sri lankans bmc res notes 2020;13(1):34
42. Becer e, mehmetcik g,
bareke h, serakinci n. Association of leptin receptor gene q223r polymorphism
on lipid profiles in comparison study between obese and non-obese subjects gene
2013;529(1):16-20.
43. Vuletic ms, milosevic vs,
jancic sa, zujovic jt, krstic ms, vukmirovic fc. Clinical significance of
leptin receptor (lepr) and endoglin (cd105) expressions in colorectal
Adenocarcinoma. J 2019;24(6):2448-2457.
44. Sarhangi n, sharifi f,
hashemian l, hassani doabsari m, heshmatzad k, rahbaran m et al. Pparg
(pro12ala) genetic variant and risk of t2dm: a systematic review and
meta-analysis sci rep 2020;29;10(1):12764.
45. Knowler wc, barrett-connor
e, fowler se, hamman rf, lachin jm, walker ea et al. Diabetes prevention
program research group. Reduction in the incidence of type 2 diabetes with
lifestyle intervention or metformin. N engl j med 2002;346(6):393-403.
46. Moazzam-jazi m,
najd-hassan-bonab l, masjoudi s. Et al. Risk of type 2 diabetes and kcnj11 gene
polymorphisms: a nested case–control study and meta-analysis sci rep
2022;12(1):20709
47. Khan v, bhatt d, khan s,
verma ak, hasan r, rafat s et al. Association of kcnj11 genetic variations with
risk of type 2 diabetes mellitus (t2dm) in north indianpopulation preprints
2019;2019070089.
48. Islam m. S.,
stimulus-secretion coupling in beta-cells: from basic to bedside, advances in
experimental medicine and biology 2020;1131;943-963.
49. Riveline jp, rousseau e,
reznik y, fetita s, philippe j, dechaume a et al. Clinical and metabolic
features of adult-onset diabetes caused by abcc8 mutations, diabetes care
2012;35(2):248-251.
50. Donath x, saint-martin c,
dubois-laforgue d, rajasingham r, mifsud f, ciangura c et al. Next-generation
sequencing identifies monogenic diabetes in 16% of patients with late
adolescence/adult-onset diabetes selected on a clinical basis: a
cross-sectional analysis, bmc medicine 2019;17(1):132.
51. Farooqi is, keogh jm, yeo
gs, et al. Clinical spectrum of obesity and mutations in the melanocortin 4
receptor gene. N engl j med 2003;348(12):1085-1095.
52. Huri hz, min ys, pendek r.
Episodes of hypoglycemia and hyperglycemia during the use of sliding scale
insulin in hospitalized diabetes patients. Asian biomed 2009;1(3):307-311.
53. Yousef aa, behiry eg, allah
wma, hussien am, abdelmoneam aa, imam mh et al. Irs-1 genetic polymorphism
(r.2963g>a) in type 2 diabetes mellitus patients associated with insulin
Resistance. Appl clin genet. 2018;28 (11):99-106.
54. Alharbi kk, khan ia,
abotalib z, al-hakeem mm. Insulin receptor substrate-1 (irs-1) gly927arg:
correlation with gestational diabetes mellitus in saudi women biomed res int
2014;146495
146499.
55. Arikoglu h, hepdogru ma,
kaya de, asik a, ipekci sh, and isciogluc f. Irs1 gene polymorphisms gly972arg
and ala513pro are not associated with insulin resistance and type 2 diabetes
risk in non-obese turkish population meta gene 2014;2:579-585.
56. Bedair rn, magour gm, ooda
sa, amar em and awad am. Insulin receptor substrate-1 g972r single nucleotide
polymorphism in egyptian patients with chronic hepatitis c virus infection
And type 2 diabetes mellitus. Egypt liver journal 2021;11(2).
57. Yadav a, kataria ma, saini
v, yadav a. Role of leptin and adiponectin in insulin resistance. Clinica
chimica acta 2013;417:80-84.
58. Nannipieri m, posadas r,
bonotti a, williams k, gonzalez-villalpando c, stern mp. Et al. Polymorphism of
the 3'-untranslated region of the leptin receptor gene, but not the adiponectin
Snp45 polymorphism, predicts type 2 diabetes: a population-based study
diabetes care 2006;29(11):2509-2511.
59. Saxena m, srivastava n,
banerjee m. Genetic association of adiponectin gene polymorphisms (+45t/g and
+10211t/g) with type 2 diabetes in north indians. Diabetes metab syndr
2012;6(2):65-69.
60. De luis da, izaola o, primo
d and aller r. Relation of a variant in adiponectin gene (rs266729) with
metabolic syndrome and diabetes mellitus type 2 in adult obese subjects,
european
Review for medical and pharmacological sciences
2020;24(20):10646-10652.
61. Guzman-ornelas mo,
chavarria-avila e, munoz-valle jf, armas-ramos le, castro-albarran j, aguilar
aldrete me et al. Association of adipoq +45t>g polymorphism with body fat
mass and blood levels of soluble adiponectin and inflammation markers in a
mexican-mestizo population, diabetes, metabolic syndrome and obesity: targets
and therapy 2012;5:369-378.