Machine Learning for Everyone: Simplifying Healthcare Analytics with BigQuery ML
Abstract
Machine learning
(ML) transforms healthcare by enabling predictive analytics, personalized
treatments and improved patient outcomes. However, traditional ML workflows
often require specialized skills, infrastructure and resources, limiting
accessibility for many healthcare professionals. This paper explores how
BigQuery ML Cloud service helps healthcare researchers and data analysts to
build and deploy models using SQL, without need for advanced ML knowledge. Our
results demonstrate that the Boosted Tree model achieved the highest
performance among the three models making it highly effective for diabetes
prediction. BigQuery ML directly integrates predictive analytics into their
workflows to inform decision-making and support patient care. We reveal this capability
through a case study on diabetes prediction using the Diabetes Health
Indicators Dataset. Our study underscores BigQuery ML’s role in democratizing
machine learning, enabling faster, scalable and efficient predictive analytics
that can directly enhance healthcare decision-making processes. This study aims
to bridge the gap between advanced machine learning and practical healthcare
analytics by providing detailed insights into BigQuery ML’s capabilities. By
demonstrating its utility in a real-world case study, we highlight its
potential to simplify complex workflows and expand access to predictive tools
for a broader audience of healthcare professionals.
Keywords: BigQuery ML;
predictive analytics; healthcare data; diabetes prediction; SQL for machine
learning; healthcare analytics; healthcare data
1. Introduction
Artificial
intelligence (AI) and machine learning (ML) are shaping the future of
healthcare by applying advanced data analysis, predictive modeling and decision
support systems. The application of AI in healthcare allows for the
identification of complex patterns in patient data, improving diagnostic
accuracy, treatment personalization and operational efficiency1. Healthcare
providers are increasingly leveraging predictive analytics to foresee health
outcomes, enabling earlier interventions and more targeted care2,26. For instance,
AI models have proven effective in identifying high-risk patients and
optimizing preventive care strategies3. Diabetes, a
major global health challenge, requires early detection and preventive care.
Predictive models built using accessible tools like BigQuery ML can help
healthcare professionals identify at-risk individuals efficiently.
Cloud computing
serves as a critical tool for AI and ML in healthcare, addressing many of the
technical and infrastructural challenges associated with large-scale data
analysis. With scalable infrastructure, cloud platforms allow healthcare
providers to process and store vast amounts of data, facilitating AI-driven
insights without the need of extensive on-site resources4. Cloud computing
reforms healthcare by reducing costs, enabling scalability, improving patient
access through telemedicine and enhancing data driven decisions. It fosters
collaboration with seamless data sharing, streamlines hospital operations and
boosts efficiency, care quality and outcomes5. Additionally,
cloud computing in healthcare offers scalability and AI integration with
addressing critical security and privacy challenges. Protecting sensitive
patient data requires implementing regulatory-compliant measures and robust
models. Research highlights various approaches to managing evolving service
models and ensuring secure and efficient healthcare systems6.
Among major cloud
providers, Amazon Web Services (AWS), Microsoft Azure and Google Cloud Platform
(GCP) offer tools for AI and ML, with high strengths. AWS provides a wide range
of machine learning tools and services such as SageMaker, which enables rapid
model development and deployment. Microsoft Azure’s AI services, including
Azure Machine Learning, are known for their integration with the broader
Microsoft ecosystem, which can be advantageous for healthcare organizations
used for Microsoft products7. Google Cloud Platform
focuses on data and machine learning services and is a powerful data warehouse
simplifying the machine learning process of users familiar with data analytics8.
Despite
advancements in AI and ML, the need for user-friendly tools that simplify
complex workflows remains critical. Traditional machine learning methods often
require extensive coding skills, which limits their accessibility to a broader
audience of healthcare practitioners. Cloud-based ML platforms provide
user-friendly tools that reduce the need for specialized expertise and
technical resources9. Through this integration,
healthcare organizations can develop, deploy and scale ML models with minimal
local infrastructure, enhancing their ability to derive information from data10.
The novelty of
this study lies in demonstrating the use of BigQuery ML to enable healthcare
professionals without programming expertise to build, train and evaluate
machine learning models directly through SQL queries. Unlike existing
cloud-based platforms, BigQuery ML simplifies the integration of predictive
analytics into healthcare workflows by eliminating the need for complex
programming or data movement. This approach has not been explored in depth in
prior research, which typically focuses on more programming-intensive platforms
such as AWS Sage Maker or Python-based workflows. The objective of this study
is to demonstrate how BigQuery ML simplifies machine learning workflows in
healthcare by enabling users to leverage SQL instead of traditional
programming-based approaches. This approach reduces technical barriers,
allowing healthcare professionals with limited programming expertise to utilize
advanced ml capabilities for decision-making and predictive analytics.
This paper is
structured as follows: Section 2 provides background on cloud-based ML
solutions and their role in healthcare analytics. Section 3 (Methodology)
describes the use of BigQuery ML for diabetes prediction, followed by a
discussion on data preparation and preprocessing techniques. Model creation,
training and hyperparameter tuning in BigQuery ML are then presented, along
with an evaluation of model performance using standard machine learning
metrics. Section 4 (Results and Discussion) highlights key findings, including
the strengths, limitations and potential applications of BigQuery ML in
healthcare. Section 5 (Future Directions and Usability Evaluation) discusses
possible extensions of this study, including real-time predictive analytics and
broader applications in healthcare. Finally, Section 6 (Conclusion) summarizes
the key contributions and outlines future research directions.
Through this structure, we aim to highlight how BigQuery ML democratizes machine learning in healthcare, making predictive analytics more accessible, scalable and efficient for a broader range of professionals.
2. Background
The intersection
of AI and healthcare has been extensively studied, with particular emphasis on
predictive analytics for chronic disease management. Diabetes, a global health
challenge affecting millions, has been a focal point for such research due to its
significant health and economic impacts. Numerous studies have demonstrated the
potential of ML models in identifying diabetes risk using health indicators
such as age, BMI and blood pressure11-13. However,
traditional ML approaches, including logistic regression and support vector
machines, often require significant computational resources and specialized
expertise, limiting their application in resource-constrained clinical
environments.
Studies indicate
that cloud-based ML has streamlined healthcare analytics by facilitating
efficient data processing and model management14. While AWS and
Azure provide comprehensive tools for machine learning, they require
significant programming expertise and do not natively integrate SQL-based
workflows. BigQuery ML, on the other hand, allows users to execute machine
learning tasks within a single environment using SQL, making it particularly
accessible for healthcare professionals without technical expertise. Table 1
summarizes the key features, advantages and limitations of BigQuery ML compared
to other leading platforms such as AWS SageMaker and Azure ML, highlighting its
unique suitability for healthcare analytics (Table 1).
Table 1. Comparison of Cloud-Based
Machine Learning Platforms: Key features, advantages and limitations of
BigQuery ML, AWS SageMaker and Azure ML, with a focus on accessibility,
integration and suitability for healthcare analytics.
|
Service |
Ease of Use |
Integration |
Expertise |
Workflow |
SQL Support |
Scalability |
Realtime |
Healthcare Use |
|
BigQuery ML |
High (SQL)
|
Google
Cloud |
Low (SQL) |
Simple |
Yes |
High |
Yes |
Strong |
|
AWS SageMaker |
Moderate
(Python) |
AWS (S3,
Lambda) |
High
(Python) |
Complex |
No |
High |
Yes |
Widely
used |
|
Azure ML |
Moderate
(GUI/Python) |
Azure
(Synapse) |
Moderate |
Mixed |
No |
High |
Yes |
Less
accessible |
Platforms like
BigQuery ML allow users to build and deploy machine learning models directly
through SQL, reducing the technical barriers associated with traditional ML
pipelines. BigQuery ML’s ability to integrate seamlessly with healthcare
datasets offers a powerful tool for healthcare professionals without
programming expertise, democratizing access to advanced analytics. Despite
these advantages, there is limited literature on the practical application of
BigQuery ML in healthcare settings, particularly for largescale datasets8. This study
leverages the Diabetes Health Indicators Dataset to demonstrate the
capabilities of BigQuery ML in a healthcare context. By utilizing SQL-based ML
workflows, we aim to provide a framework for healthcare professionals to
implement predictive analytics with minimal technical overhead, thereby
addressing a critical gap in existing research.
2.1.
Related studies and literature gap
Most existing
research relies on programming-intensive platforms requiring Python, R or
specialized ML frameworks, which create barriers for non-technical users.
Limited studies explore SQL-based machine learning tools like BigQuery ML,
which provide an accessible alternative for healthcare professionals lacking
programming expertise. While studies have evaluated ML performance in
healthcare using conventional programming methods, the accessibility and
efficiency of SQL-driven ML workflows remain underexplored.
This gap is
particularly significant given that SQL is widely used in healthcare data
analytics27-29, making it a natural choice for enabling predictive
modeling without additional programming knowledge. Prior studies have not
focused on how healthcare professionals can leverage SQL for ML without
requiring external tools, manual data movement or complex ML frameworks.
BigQuery ML addresses this gap by allowing professionals to leverage the power
of machine learning directly through SQL commands, making it an ideal solution
for real-world healthcare applications. SQL remains one of the most widely used
programming languages worldwide, consistently ranking among the top three in
2024 [27]. This widespread familiarity makes SQL a natural choice for professionals
in healthcare analytics, enabling them to leverage existing skills for advanced
machine learning tasks without requiring extensive technical training.
While alternative
tools like Weka and ChatGPT offer unique approaches to machine learning
workflows, they fall short in key aspects that are critical for real-world
healthcare applications. Weka, for example, is a GUI-driven tool suitable for
standalone ML studies but lacks integration with cloud platforms, making it
inefficient for handling large-scale datasets or real-time analysis.
Additionally, Weka requires users to export and import data manually,
increasing the likelihood of errors and inefficiencies. Similarly, while
ChatGPT can generate ML code through descriptive prompts, it assumes the user
has a basic understanding of programming to interpret, test and refine the
code. Furthermore, ChatGPT does not provide a cloud environment to execute the
generated code or access integrated tools such as BigQuery, Cloud Storage or
Vertex AI. In contrast, BigQuery ML operates within the Google Cloud ecosystem,
enabling direct interaction with databases and other cloud tools. This
capability streamlines the entire data workflow, from ingestion to model
training and deployment, without requiring external environments or tools.
This study addresses this gap by showcasing BigQuery ML as a fully integrated cloud-based ML solution, demonstrating how it can be applied to structured healthcare data without requiring programming expertise or ML-specific knowledge.
2.2.
Research question
Given the
increasing demand for accessible ML solutions in healthcare and the gap in
research on SQLdriven ML platforms, this study aims to address the following
research question:
"How
effectively can BigQuery ML be utilized for predictive healthcare analytics and
how does it compare in terms of accessibility, ease of use and performance to
other cloud-based ML solutions?"
This question is
driven by the need to evaluate the feasibility of SQL-based ML workflows for
healthcare professionals, as well as the potential advantages and limitations
of using BigQuery ML for predictive analytics. By investigating this question,
the study seeks to determine whether BigQuery ML can lower technical barriers
to machine learning adoption in healthcare, providing a scalable, efficient and
accessible alternative to programming-heavy ML platforms.
3. Methodology
This section
describes the end-to-end workflow for implementing machine learning models
using BigQuery ML within Google Cloud Platform (GCP) for healthcare analytics.
We first introduce the cloud infrastructure used for data storage, processing
and model execution, followed by an explanation of the BigQuery data warehouse
and its role in handling large-scale structured healthcare datasets. Next, we
outline the ML models employed-Logistic Regression, Boosted Tree and Deep
Neural Network (DNN)- and justify their selection for this study. Additionally,
we provide details about the dataset, including its characteristics,
preprocessing considerations and rationale for use. The section concludes with
a discussion on hyperparameter tuning and model evaluation (Figure 1).
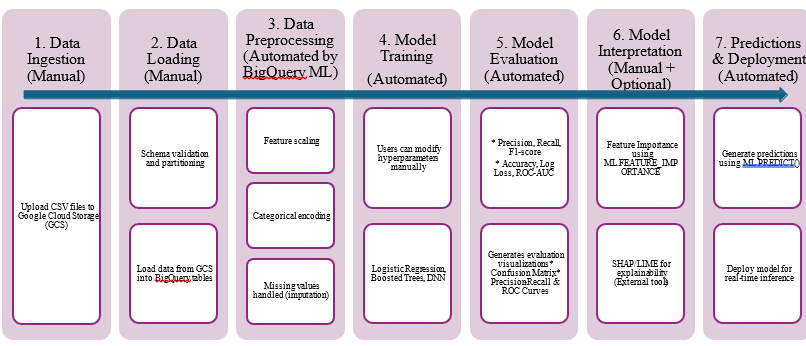
Figure 1: Workflow of BigQuery ML-based machine learning pipeline.
3.1.
Work environment
Google Cloud
Platform (GCP) provides a robust, scalable cloud infrastructure for data
storage, processing and ML analysis. Its suite of tools enables end-to-end data
workflows, from ingestion to predictive modeling, while ensuring compliance
with healthcare regulations such as HIPAA [15]. The Google Cloud ecosystem,
including BigQuery, Cloud Storage and BigQuery ML, was chosen for this study
due to its efficiency in handling large-scale healthcare datasets securely and
seamlessly16.
BigQuery, a fully
managed serverless data warehouse, was utilized for data storage, querying and
preprocessing. Its low-latency SQL-based architecture is particularly
well-suited for healthcare analytics, allowing rapid data exploration without
the need for additional infrastructure management17. By storing the
Diabetes Health Indicators Dataset in BigQuery, we leveraged its built-in
preprocessing capabilities while maintaining data security and integrity.
BigQuery ML was
used to build, train and evaluate ML models directly through SQL queries. This
SQL based approach democratizes ML access, allowing healthcare professionals
without programming expertise to implement predictive analytics with minimal
overhead18.
Hyperparameter tuning was conducted through iterative SQL-based adjustments in BigQuery ML. Learning rate, maximum iterations and tree depth were optimized using ML. EVALUATE functions, ensuring a balance between precision and recall for healthcare predictions. Since BigQuery ML operates as a fully managed cloud service, no on-premise hardware was required for model execution. The computational workload was handled by Google Cloud’s distributed infrastructure, leveraging high-performance computing resources optimized for SQL-based ML processing. This eliminates the need for dedicated GPUs or TPUs typically required for deep learning tasks, reducing hardware dependencies while ensuring scalability for large datasets.
3.2.
Dataset used and rationale
The Behavioral
Risk Factor Surveillance System (BRFSS) is an annual health survey conducted by
the CDC since 1984. This telephone survey collects health-related data from
over 400,000 Americans each year, focusing on health risk behaviors, chronic
conditions and preventive healthcare services. For this study, we used a
refined 2015 BRFSS dataset, publicly available in CSV format on Kaggle. The
dataset includes responses from 70,692 individuals, evenly balanced between
non-diabetic (0) and pre-diabetic or diabetic (1) individuals. The target
variable for this study is Diabetes binary, which indicates whether a
respondent has diabetes (1) or not (0).
To build our predictive models, we selected 21 key health indicators relevant to diabetes prediction. These features include demographic factors, lifestyle habits and clinical indicators that have been commonly linked to diabetes risk in prior research22. (Table 2) provides a summary of the 21 selected features, their type (numerical/categorical), key statistics and brief descriptions:
Table 2:
Summary of Selected Features in the Dataset.
Feature
Description
Type
High BP
High blood
pressure (1 = Yes, 0 = No)
Categorical
High Chol
High
cholesterol (1 = Yes, 0 = No)
Categorical
Chol Check
Cholesterol
check in past 5 years (1 = Yes, 0 = No)
Categorical
BMI
Body Mass
Index
Numerical
Smoker
Smoked at
least 100 cigarettes in lifetime (1 = Yes, 0 = No)
Categorical
Stroke
History of
stroke (1 = Yes, 0 = No)
Categorical
Heart Disease or Attack
Coronary
heart disease or heart attack (1 = Yes, 0 = No)
Categorical
Phys Activity
Engages in
physical activity outside work (1 = Yes, 0 = No)
Categorical
Fruits
Consumes
fruit daily (1 = Yes, 0 = No)
Categorical
Veggies
Consumes
vegetables daily (1 = Yes, 0 = No)
Categorical
HvyAlcoholConsump
Heavy
alcohol consumption (1 = Yes, 0 = No)
Categorical
AnyHealthcare
Has
healthcare coverage (1 = Yes, 0 = No)
Categorical
NoDocbcCost
Couldn’t
see doctor due to cost (1 = Yes, 0 = No)
Categorical
GenHlth
General
health rating (1 = Excellent, 5 = Poor)
Ordinal
MentHlth
Days of
poor mental health in past month (0–30)
Numerical
PhysHlth
Days of
poor physical health in past month (0–30)
Numerical
DiffWalk
Difficulty
walking or climbing stairs (1 = Yes, 0 = No)
Categorical
Feature
Description
Type
High BP
High blood
pressure (1 = Yes, 0 = No)
Categorical
High Chol
High
cholesterol (1 = Yes, 0 = No)
Categorical
Chol Check
Cholesterol
check in past 5 years (1 = Yes, 0 = No)
Categorical
BMI
Body Mass
Index
Numerical
Smoker
Smoked at
least 100 cigarettes in lifetime (1 = Yes, 0 = No)
Categorical
Stroke
History of
stroke (1 = Yes, 0 = No)
Categorical
Heart Disease or Attack
Coronary
heart disease or heart attack (1 = Yes, 0 = No)
Categorical
Phys Activity
Engages in
physical activity outside work (1 = Yes, 0 = No)
Categorical
Fruits
Consumes
fruit daily (1 = Yes, 0 = No)
Categorical
Veggies
Consumes
vegetables daily (1 = Yes, 0 = No)
Categorical
HvyAlcoholConsump
Heavy
alcohol consumption (1 = Yes, 0 = No)
Categorical
AnyHealthcare
Has
healthcare coverage (1 = Yes, 0 = No)
Categorical
NoDocbcCost
Couldn’t
see doctor due to cost (1 = Yes, 0 = No)
Categorical
GenHlth
General
health rating (1 = Excellent, 5 = Poor)
Ordinal
MentHlth
Days of
poor mental health in past month (0–30)
Numerical
PhysHlth
Days of
poor physical health in past month (0–30)
Numerical
DiffWalk
Difficulty
walking or climbing stairs (1 = Yes, 0 = No)
Categorical
Sex
Gender (1 = Male,
0 = Female) Categorical
|
Age |
Age category (1
= 18-24, 13 = 80+) |
Ordinal |
|
Education |
Education level
(1 = No school, 6 = College graduate) |
Ordinal |
|
Income |
Income category
(1 = < $10,000, 8 = > $75,000) |
Ordinal |
|
Target |
Description |
Type |
|
Diabetes binary |
Diabetes status
(1 = Diabetic/Pre-diabetic, 0 = Nondiabetic) |
Categorical |
3.3.
Machine learning models
To apply BigQuery
ML for healthcare analytics, three predictive models were selected: Logistic
Regression, Boosted Trees and Deep Neural Networks (DNN). While BigQuery ML
supports a range of models, these models span a spectrum of complexity, from
interpretable linear models to advanced ensemble methods and deep learning
architectures. Their selection is based on prior research in healthcare
predictive modeling, where they have shown effectiveness in disease prediction
tasks.
· Logistic regression: Chosen as a baseline model
due to its interpretability and effectiveness for binary classification, making
it particularly useful in clinical decision-making19.
· Boosted trees: Selected for its ability to
capture complex relationships between features and improve classification
accuracy through an iterative learning process20.
· Deep Neural Networks (DNN): Applied to
explore the potential of deep learning in detecting high dimensional patterns
within structured healthcare datasets21.
Justification for
model selection is that while BigQuery ML supports various machine learning
models, these three were deliberately chosen to showcase its capabilities
across different levels of complexity; Logistic Regression serves as an
interpretable, easy-to-use model for healthcare professionals with minimal ML
expertise. Boosted Trees provide an intermediate-level approach, offering
improved predictive performance while remaining interpretable. Deep Neural
Networks (DNNs) demonstrate how BigQuery ML can handle more complex,
computationally demanding models without requiring external tools or custom
infrastructure.
This selection
highlights how BigQuery ML enables users to scale from simple ML applications
to advanced AI-driven healthcare solutions within a SQL-based environment.
Future work can explore additional models, but this study focuses on
demonstrating BigQuery ML’s ease of use and versatility in healthcare
analytics.
3.4. Data loading
The Diabetes Health Indicators Dataset was first uploaded from Google
Cloud Storage (GCS) to BigQuery. This process involved multiple steps to ensure
the data was correctly ingested and prepared for machine learning tasks. The
following outlines the key stages:
3.4.1. Uploading the dataset
to google cloud storage:
The raw dataset, diabetes_binary_health_indicators_BRFSS2015.csv,
was stored as a CSV file in an existing or newly created GCS bucket. This setup
facilitated easy access for data ingestion into BigQuery, ensuring data
security and scalability.
3.4.2. Creating the big
query dataset: A new dataset named diabetes_analysis was created in BigQuery to store the data.
Configuration details such as Dataset ID and Data Location were specified
according to project requirements. This step established a structured
repository for managing the dataset within BigQuery.
3.4.3. Loading data from
GCS to BigQuery: The CSV data was loaded into BigQuery by
creating a table within the diabetes analysis dataset. Configuration settings
included:
Source: GCS path
(e.g.,
gs://your_bucket/diabetes_binary_5050split_health_indicators_BRFSS2015.csv).
File Format: CSV
Destination Table: diabetes data.
BigQuery ML automatically partitions the data into training and evaluation sets during model training using parameters such as data_split_method and data_split_eval_fraction. This eliminates the need for manual data splitting and ensures a standardized approach to model evaluation. Additionally, BigQuery ML validates the schema to ensure compatibility during ingestion and training, reducing the likelihood of errors caused by data mismatches.
3.5.
Data preparation
3.5.1. Schema validation
and compatibility: BigQuery ML validates
the schema during ingestion and training to ensure compatibility between the
dataset and the machine learning model. This step reduces the likelihood of
errors caused by data mismatches, streamlining the overall process and maintaining
data integrity. By leveraging Google Cloud Storage and BigQuery, the data
preparation and loading process became highly efficient and scalable, laying a
strong foundation for subsequent model creation and training tasks.
3.5.2. Automatic
preprocessing in BigQuery ML: One of the
advantages of BigQuery ML is its built-in automatic preprocessing, which
eliminates the need for extensive manual data preparation. Automatic
preprocessing consists of missing value imputation and feature transformations,
ensuring consistency and efficiency in model training.
3.5.3. Missing value
imputation: BigQuery ML automatically imputes missing
values based on data type. For example, numerical features are replaced with
the mean value, while categorical features are assigned to a special missing
category. This prevents model failures due to missing data and ensures robust
training.
3.5.4. Feature
transformations: Feature Transformations: BigQuery ML
automatically standardizes numerical features (zero mean, unit variance) for
logistic regression and DNNs, ensuring consistent feature scaling. Boosted
trees and random forests do not require standardization, as they handle raw
feature values natively.
3.5.5. Categorical
feature encoding: BigQuery ML supports
various encoding methods for categorical variables, including one-hot encoding,
dummy encoding, label encoding and target encoding. The choice of encoding
depends on the model type and details are available in the official
documentation30.
3.5.6. Manual data
preprocessing using SQL: While BigQuery ML
automates preprocessing, additional SQL based feature engineering techniques
can be applied for greater customization. Missing values can be handled with
SQL queries by applying imputation techniques-numerical variables can be filled
using the median and categorical variables can be encoded using one-hot
encoding. Additionally, feature selection can be conducted by calculating
correlations between each feature and the target variable (diabetes_binary)
using SQL-based statistical functions. These features streamline the data
preparation process, allowing all preprocessing steps to be completed within
the BigQuery environment without requiring external tools or programming. (Figure
2.a) demonstrates a sample SQL query for handling missing values, while (Figure
2.b) shows an example of categorical encoding. (Figure 2.c)
illustrates how SQL can be used to calculate correlations for feature
selection.
|
|
|
|
|
|
|
|
|
|
Figure 2.a: Handling missing values by
applying median imputation using SQL example.
|
|
Figure 2.b: SQL-based one-hot encoding
of categorical variables example.
Figure 2.c: Calculating feature
correlation with the target variable using SQL example.
3.5.
Model creation and training
We employed BigQuery ML to train three models: Logistic Regression,
Boosted Tree and Deep Neural Network (DNN), leveraging SQL for all training
workflows. BigQuery ML enables healthcare professionals to build machine
learning models using SQL commands, simplifying the process for users without
programming expertise. Each model was trained on the Diabetes Health Indicators
Dataset with 21 feature variables.
We explain the SQL query for the Boosted Tree model in this section as
an example. The SQL query used to create and train the Boosted Tree model is
shown in (Figure 3). This query demonstrates the simplicity and
flexibility of BigQuery ML’s SQL-based approach for model creation.
|
|
Figure 3. SQL query used for creating
and training the Boosted Tree model in BigQuery ML. The query includes key
hyperparameters such as learning rate, maximum iterations and regularization
terms to optimize model performance.
Here is a
line-by-line explanation of the SQL query:
· CREATE OR REPLACE
MODEL: This command creates a new machine learning model or replaces an
existing one with the same name. The model is stored in the specified project
and dataset under the name diabetes_model.
· OPTIONS Clause: This section
specifies the configuration settings for the Boosted Tree model: model_type = 'boosted_tree_classifier':
Indicates that the model is a Boosted Tree classifier, designed for binary
classification tasks. input_label_cols =
['Diabetes_binary']: Specifies that the target column for prediction is
Diabetes_binary. data_split_method =
'RANDOM': Instructs BigQuery ML to randomly split the data into training
and evaluation sets. data_split_eval_fraction
= 0.2: Allocates 20% of the data for evaluation and 80% for training. max_iterations = 150: Sets the maximum
number of iterations for the boosting process to converge. learn_rate = 0.05: Defines the learning rate, which controls the
contribution of each tree to the final prediction. min_rel_progress = 0.00001: Specifies the minimum relative progress
required between iterations to avoid early stopping. l1_reg = 0.1: Applies L1 regularization to the model to prevent
overfitting by penalizing large coefficients. l2_reg = 2.0: Applies L2 regularization, adding an additional
penalty for large coefficients to enhance generalization.
· AS SELECT Clause:
o SELECT *: Selects all
columns from the dataset for model training, ensuring that all relevant
features and the target variable are included.
o FROM
'project_id.dataset_id.diabetes_data': Specifies the source table
containing the dataset.
o WHERE Diabetes_binary IS NOT
NULL: Filters out rows with null values in the target column
(Diabetes_binary), ensuring clean data for training.
The Boosted Tree
model was designed to handle complex relationships in the data, leveraging
BigQuery ML’s efficient infrastructure. The use of hyperparameters such as
max_iterations, learn_rate and regularization settings (L1 and L2) ensures the
model achieves a balance between accuracy and generalization, critical for
healthcare applications like diabetes prediction. This SQL-based approach
demonstrates the accessibility of advanced machine learning methods for
healthcare professionals without the need for programming expertise.
Hyperparameter tuning in BigQuery ML was conducted using iterative adjustments
to model parameters through the OPTIONS clause in SQL queries. For example, for
the Boosted Tree model, parameters such as learning rate, max iterations and
tree depth were specified and adjusted iteratively. Each model was evaluated
using the ML.EVALUATE function to identify the optimal settings based on
metrics such as log loss and F1 score. This approach ensured that the models
achieved a balance between precision and recall, critical for healthcare
predictions.
4. Results and Discussion
This section
evaluates the performance of Logistic Regression, Boosted Tree and Deep Neural
Network (DNN) models using key metrics like precision, recall, F1 score, ROC
AUC, accuracy and log loss. Section 4.2 explains these metrics and their
relevance in healthcare analytics. Section 4.3 examines confidence threshold
trade-offs, while Section 4.4 analyzes precision-recall curves. Section 4.5
discusses ROC curves and model discrimination ability. Section 4.6 provides a
comparative analysis of model performance, summarized in Table 1. Finally,
Section 4.7 explores practical implications, guiding model selection for
healthcare applications.
4.1. Evaluation metrics
Model evaluation
is a critical aspect of the machine learning process, especially in healthcare
applications where predictive accuracy has real-world implications. The
performance of the models, Logistic Regression, Boosted Tree and Deep Neural
Network (DNN), was assessed using multiple metrics, including precision,
recall, F1 score, ROC AUC, accuracy and log loss. BigQuery ML provides these
metrics along with visual tools such as confidence threshold plots,
precision-recall curves and ROC curves. These tools are particularly useful in
healthcare contexts for exploring model performance across various thresholds
and understanding trade-offs between sensitivity and specificity, offering
deeper insights into the strengths and limitations of each model.
In BigQuery ML,
each trained model is automatically saved within the specified dataset. This
built-in storage allows users to retrieve performance metrics directly from the
model and streamlines the evaluation process. BigQuery ML’s ML. PREDICT
function also allows users to generate predictions on new or unseen data
directly within BigQuery, integrating predictive analytics seamlessly into
healthcare workflows.
The key
evaluation metrics used in this study include:
4.1.1. Precision (P): Measures the proportion of correctly predicted positive cases among
all positive predictions. High precision reduces false positives23.
??
????????? = ?? + ??
where TP is true positives and FP is false positive.
4.1.2. Recall (Sensitivity, R): Represents the proportion of actual positive cases correctly
identified by the model. High recall ensures minimal false negatives23.
??
?? + ??
where FN is false negatives.
4.1.3. F1 score: The harmonic means of
precision and recall, balancing the two metrics24,25.
????????? × ??????
????????? + ??????
4.1.4. Accuracy (A): The ratio of
correctly predicted cases (both positive and negative) to the total cases.
?? + ??
?? + ?? + ?? + ??
where TN is true negative.
4.1.5. Log Loss: A metric used for
probabilistic predictions, penalizing incorrect confidence levels. Lower log
loss indicates better performance.
?
??? ????
?
N represents the
total number of samples, ?? denotes the
actual class label (0 or 1) and ?̂? refers to the
predicted probability of the positive class (1).
4.1.6. Receiver Operating Characteristic (ROC) AUC:
Measures the ability of the model to distinguish between classes across
different thresholds. A higher AUC indicates better discrimination between
positive and negative cases.
???
These metrics
provide a comprehensive evaluation of the models, ensuring a balanced
assessment of their predictive performance in healthcare applications.
4.2.
Confidence threshold vs precision/recall
The leftmost
panels in Figure 4 depict the relationship between confidence thresholds,
precision and recall for the three models. These graphs illustrate how the
precision and recall values change as the confidence threshold is adjusted.
Precision measures the proportion of true positive predictions among all
positive predictions, while recall indicates the proportion of actual positive
cases correctly identified by the model.
For the Boosted
Tree model, precision and recall intersect at a balanced threshold,
highlighting its ability to maintain a robust trade-off between the two
metrics. The DNN model exhibits a similar pattern, with both precision and
recall values demonstrating consistent performance as the threshold varies. In
contrast, the Logistic Regression model shows a steeper decline in recall as
the threshold increases, emphasizing its limitations in preserving sensitivity
at higher confidence levels.
Understanding
these trade-offs is crucial in healthcare, where high recall is often
prioritized to minimize false negatives, such as undiagnosed diabetes cases. By
analyzing these graphs, practitioners can identify thresholds that align with
clinical priorities, ensuring that the selected model meets the specific needs
of the application while balancing precision and recall effectively (Figure
4).
a)
b)
c)
4.3.
Precision-recall curve
The middle panels in Figure 4 illustrate the precision-recall curves for the models. These curves provide a detailed view of the relationship between precision and recall across different thresholds. The area under the precision-recall curve (AUC-PR) is a critical metric for evaluating a model's ability to maintain high recall without sacrificing precision. For the Boosted Tree model, the AUC-PR is 0.808, indicating strong performance in identifying true positives while minimizing false positives. This robustness is particularly valuable in healthcare applications, where accurate identification of diabetes cases is essential. The DNN model achieved an AUC-PR of 0.797, slightly lower than the Boosted Tree but still demonstrates reliable performance. In contrast, the Logistic Regression model exhibited an AUC-PR of 0.748, highlighting its comparatively lower ability to balance precision and recall. Precision-recall curves are especially useful for imbalanced datasets, such as the diabetes dataset, where the positive class (diabetes cases) is relatively rare. By analyzing these curves, healthcare practitioners can gain insights into the models' trade-offs, enabling the selection of the most suitable model for their specific needs.
4.4.
Receiver Operating Characteristic (ROC) curve
The rightmost panels in Figure 4 show the Receiver Operating Characteristic (ROC) curves for the models. These curves are essential for evaluating a model's ability to distinguish between positive and negative classes by plotting the true positive rate (recall) against the false positive rate across various thresholds. The area under the ROC curve (AUC-ROC) provides a single, comprehensive metric to summarize the model’s discriminatory power. For the Boosted Tree model, the AUC-ROC is 0.834, demonstrating its strong ability to differentiate between individuals with and without diabetes. This high value highlights the model's reliability in achieving a balance between sensitivity and specificity, critical for healthcare applications. The DNN model follows closely with an AUC-ROC of 0.83, showcasing comparable discriminatory power. The Logistic Regression model, while simpler, achieved an AUC-ROC of 0.778, indicating a modest but effective performance in distinguishing between the two classes. ROC curves are particularly valuable in healthcare scenarios were minimizing false negatives (e.g., missed diabetes cases) is essential. By analyzing the ROC curves, healthcare professionals can determine optimal thresholds that prioritize sensitivity while maintaining reasonable specificity, ensuring effective interventions for at-risk patients.
4.5.
Comparative insights
Among the evaluated models, the Boosted Tree model achieved the highest
performance, with an F1-score of 0.7626 and an AUC-ROC of 0.8339 (see Table 1).
These metrics were chosen to determine the best performing model as they
provide a balanced assessment of classification performance, particularly in
healthcare applications where both false positives and false negatives must be
minimized. The F1-score is widely recognized as an essential metric when dealing
with class imbalance, as it accounts for both precision and recall. In diabetes
prediction, missing a positive case (false negative) can lead to delayed
diagnosis and worsened health outcomes, making recall particularly critical.
However, an excessive focus on recall could lead to more false positives, which
may overburden healthcare resources. Therefore, a model that optimally balances
these factors-such as Boosted Tree-offers clinical and operational advantages
in a real-world healthcare setting.
Similarly, AUC-ROC is a crucial metric for assessing a model's ability
to distinguish between diabetic and non-diabetic patients across various
decision thresholds. A higher AUC-ROC score indicates that the model
consistently ranks positive cases above negative ones, making it particularly
valuable for risk stratification in healthcare applications. The Boosted Tree
model outperformed Logistic Regression and DNN in both F1-score and AUC-ROC,
demonstrating better generalization and superior classification performance in
this study. Based on these findings, the Boosted Tree model is recommended for
diabetes prediction using BigQuery ML (Table 3).
Table 3: Evaluation metrics of
Logistic Regression, Boosted Tree and DNN models trained in BigQuery ML.
|
|
Precision |
Recall |
Accuracy |
F1 Score |
Log Loss |
ROC AUC |
|
Logistic Regression |
0.6972 |
0.7443 |
0.708 |
0.72 |
0.5674 |
0.7766 |
|
Boosted Tree |
0.7324 |
0.7954 |
0.7546 |
0.7626 |
0.498 |
0.8339 |
|
DNN |
0.7185 |
0.8215 |
0.752 |
0.7665 |
0.5049 |
0.8295 |
4.6.
Feature importance analysis for the boosted tree model
Feature importance helps us understand which variables contribute the
most to the model’s predictions. For the boosted tree classifier, we used
importance gain as the primary metric. Importance gains measures how much a
feature improves decision splits in the model, meaning higher values indicate
more influential predictors. The results indicate that High Blood Pressure
(HighBP) is the most significant predictor of diabetes, followed by General
Health (GenHlth). This aligns with medical expectations, as high blood pressure
and overall health perception are strongly associated with metabolic disorders,
including diabetes. Other key features influencing diabetes prediction include
Age, BMI and High Cholesterol (HighChol), all of which are well-documented risk
factors for diabetes. Difficulty Walking (DiffWalk), Heavy Alcohol Consumption
and History of Heart Disease or Attack also show moderate importance,
reflecting broader cardiovascular risks associated with diabetes. Lifestyle
factors such as Physical Activity, Smoking Status and Vegetable Consumption
have relatively lower importance in this model. While these factors influence
overall health, their specific contribution to diabetes risk is smaller
compared to the dominant predictors. To better illustrate the impact of each
feature, we present a bar chart showing the importance gain of all features in
Figure 5. Higher values indicate stronger predictive power (Figure 5).
Figure 5. Feature Importance in
Boosted Tree Model for Diabetes Prediction
In conclusion,
the feature importance results align well with medical knowledge. Blood
pressure, general health, age, BMI and cholesterol levels play key roles in
predicting diabetes. These insights can help refine predictive models and
inform healthcare interventions.
4.7.
Discussion and practical implications
4.7.1
Strengths of BigQuery ML:
· Ease of use: The SQL-based
interface of BigQuery ML allows users to perform end-to-end machine learning
workflows-including data preparation, model training, evaluation and prediction-entirely
within BigQuery. This simplicity enables healthcare researchers, data analysts
and non-technical professionals to integrate predictive analytics into their
decision-making processes without requiring advanced ML expertise. BigQuery ML also serves as a bridge
for professionals transitioning into data engineering or data science roles by
offering an accessible introduction to machine learning concepts using SQL.
Non-technical managers and analysts can leverage BigQuery ML to generate
insights from healthcare data without coding in Python or other programming
languages.
· Scalability &
speed: BigQuery ML’s serverless architecture allows for the efficient
processing of largescale healthcare datasets in real time. Unlike traditional
machine learning workflows that require manual ETL pipelines, BigQuery ML
integrates data storage, preprocessing and model training into a single
environment, reducing the need for additional computational infrastructure. This scalability makes it suitable for
population-level health studies, predictive modeling for disease risk
assessment and large-scale patient data analytics. By leveraging Google Cloud’s
infrastructure, healthcare organizations can process and analyze millions of
records without significant latency. While BigQuery ML benefits from cloud
scalability organizations with cost or resource constraints may need to
optimize queries and select cost effective models to balance performance and
affordability.
· Seamless integration
with healthcare workflows: BigQuery ML’s ability to integrate directly with
Google Cloud services-such as Cloud Storage, Cloud Functions and Vertex AI-enables
a streamlined machine learning workflow for healthcare applications. This
allows automated model updates, real-time predictions and interoperability with
electronic health records (EHRs) and other healthcare platforms. By reducing technical barriers and
infrastructure overhead, BigQuery ML makes predictive healthcare analytics more
accessible to hospitals, clinics and public health organizations.
4.7.2.
Limitations and challenges:
· Dataset biases
and generalizability: While this study utilizes a balanced diabetes
dataset, healthcare datasets often suffer from biases related to demographics,
socioeconomic factors and healthcare accessibility. Since our dataset is
derived from a specific population, it may not generalize well to other groups.
To mitigate bias and improve fairness, future
research should explore external validation with diverse datasets to assess
model robustness across different populations. Additionally, bias detection
techniques and fairness-aware machine learning approaches could be integrated
into BigQuery ML workflows.
· Interpretability
of complex models: While Boosted Trees and Logistic Regression models
in BigQuery ML offer transparency and interpretability, Deep Neural Networks
(DNNs) lack explainability, making them less suitable for high-stakes medical
decision-making. To improve
interpretability, techniques such as SHAP (Shapley Additive Explanations) and
LIME (Local Interpretable Model-agnostic Explanations) can be integrated with
BigQuery ML workflows. Future research can explore post-hoc interpretability
frameworks that enhance model trustworthiness for healthcare applications.
· Image-based
machine learning limitations: BigQuery ML is optimized for structured,
tabular datasets and does not natively support image-based machine learning.
For healthcare applications involving medical imaging (e.g., X-rays, MRIs, CT
scans), specialized tools such as Google Cloud Auto ML Vision and TensorFlow
should be considered. However,
metadata associated with image datasets (e.g., diagnostic labels, patient
history) can still be processed using BigQuery ML, providing valuable insights
when combined with clinical data. Future work can explore hybrid workflows,
where image analysis is performed using deep learning models in Vertex AI or
TensorFlow, while BigQuery ML is used for structured data integration and
analysis.
· Manual hyperparameter tuning: BigQuery ML currently does not support automated hyperparameter tuning (e.g., grid search or random search). Instead, users must manually specify parameters in SQL queries, adjusting learning rates, iterations and tree depths based on model performance. This manual tuning process may require additional effort compared to platforms like AWS Sage Maker or Azure ML, which offer automated tuning features. Future versions of BigQuery ML could benefit from integrating Auto ML like hyperparameter optimization to further enhance usability.
5. Future Directions and
Usability Evaluation
This study represents one of the first investigations into cloud-based SQL-driven machine learning workflows in healthcare. Future research should expand its applicability to a wider range of medical datasets and real-time predictive healthcare systems.
5.1.
Future research directions
5.1.1. Real-time predictive analytics integration: Future work
should explore the integration of BigQuery ML into real-time healthcare
systems, such as hospital EHR systems, telemedicine platforms and public health
dashboards. Real-time analytics could enable immediate identification of
high-risk patients, allowing timely interventions and improving patient
outcomes.
5.1.2. Expanding dataset diversity: While this study
focused on diabetes prediction, future research should explore broader
healthcare challenges, such as cardiovascular diseases, cancer detection and
mental health.
Additionally,
extending BigQuery ML’s application to image-based medical datasets-such as
X-rays, MRIs and CT scans-could be an important area for further investigation.
A hybrid approach integrating BigQuery ML for structured data with deep
learning models in Vertex AI or TensorFlow for image analysis could enhance
predictive insights and broaden the scope of healthcare analytics.
5.1.3. Ethical considerations and bias mitigation: Ethical challenges, such as algorithmic bias and fairness, must be addressed in future research. Developing fairness-aware models and ensuring that BigQuery ML workflows are compliant with healthcare regulations (e.g., HIPAA) will be essential for equitable and responsible deployment.
5.2.
Possible applications of the research
5.2.1. Personalized healthcare: BigQuery ML can
support the creation of personalized treatment plans by predicting individual
risks for chronic conditions such as diabetes. High-risk patients could benefit
from tailored interventions, including dietary adjustments, exercise regimens
and medication schedules.
5.2.2. Population health management: Public health
organizations can leverage BigQuery ML to analyze population health trends,
enabling resource allocation to communities with higher health risks. For
example, targeted diabetes prevention campaigns could reduce disease prevalence
and associated healthcare costs.
5.2.3. Clinical Decision Support Systems (CDSS): Integrating
BigQuery ML predictions into CDSS could assist healthcare providers by flagging
high-risk patients during routine checkups, reducing diagnostic delays and
improving treatment outcomes.
5.2.4. Remote monitoring and telemedicine: BigQuery ML can
analyze real-time data from wearable devices or remote monitoring systems,
enabling continuous diabetes management and reducing hospital visits for
chronic patients.
5.2.5. Risk assessment for health insurance: Health insurers
can use BigQuery ML to optimize risk assessments, personalize insurance plans
and incentivize preventive care, ultimately reducing costs and improving
patient outcomes.
5.2.6. Educational tools for healthcare professionals: BigQuery ML’s SQL-based approach can serve as an educational tool for healthcare professionals, teaching them the fundamentals of predictive analytics and encouraging broader adoption of machine learning in clinical settings.
5.3.
Proposed usability evaluation
While this study
shows BigQuery ML’s capabilities, a formal usability evaluation could provide
deeper insights into its effectiveness for healthcare professionals. A
structured questionnaire comparing BigQuery ML, WEKA and Python-based tools
could assess:
· Ease of use: Comparing the SQL-based
workflow with GUI-driven (WEKA) and code-based (Python) tools.
· Tool preference: Evaluating user preference
based on familiarity, flexibility and usability.
· Cloud services: Assessing the importance of
scalability, integration and real-time analytics in ML tool selection.
· Efficiency: Measuring workflow speed
and reduction in external tool dependency.
· Overall satisfaction: Rating usability and the likelihood of recommending BigQuery ML.Such an evaluation would validate BigQuery ML’s practicality in healthcare ML workflows and justify the benefits of SQL-based machine learning tools in cloud environments.
6. Conclusion
This study
demonstrated how BigQuery ML simplifies machine learning workflows in
healthcare by enabling predictive modeling through SQL-based queries. By
eliminating the need for extensive programming expertise, BigQuery ML makes
machine learning more accessible to healthcare professionals, data analysts and
researchers. Through a case study on diabetes prediction using the Diabetes
Health Indicators Dataset, we evaluated three predictive models-Logistic
Regression, Boosted Tree and Deep Neural Networks (DNN)-and identified the
Boosted Tree model as the best-performing approach based on F1-score and ROC
AUC.
The results
confirm that BigQuery ML’s built-in preprocessing, scalability and ease of
integration with cloud-based healthcare data pipelines make it a valuable tool
for predictive analytics. The feature importance analysis revealed that High
Blood Pressure, General Health, BMI and Age are significant predictors of
diabetes, aligning with medical expectations. These findings underscore
BigQuery ML’s capability to support real-world healthcare applications,
particularly in identifying high-risk individuals for early intervention. While
BigQuery ML offers significant advantages in accessibility and efficiency,
certain challenges remain, including model interpretability, dataset biases and
the lack of automated hyperparameter tuning. Future research should explore
methods to enhance explainability, integrate real time predictive analytics
into healthcare systems and expand the application of BigQuery ML to other
diseases and medical datasets. The study highlights the broader impact of
cloud-based machine learning on healthcare analytics, paving the way for more
scalable, interpretable and data-driven decision-making. By bridging the gap
between advanced AI techniques and practical healthcare use cases, BigQuery ML
provides a cost-effective and accessible solution for predictive analytics,
supporting the future of personalized medicine and population health
management.
7. References
- Topol EJ, High-performance
medicine: the convergence of human and artificial intelligence, Nature
medicine, 2019;25: 44-56.
- Rajkomar A, Dean J, Kohane I. Machine Learning in
Medicine Reply, The New England journal of medicine, 2019;380: 2589-2590.
- Miotto Riccardo, et al. Deep learning for
healthcare: review, opportunities and challenges, Briefings in bioinformatics, 2018:
1236-1246.
- Raghupathi W, Raghupathi V.
Big data analytics in healthcare: promise and potential, Health information
science and systems, 2014;2:1.
- https://flatirons.com/blog/cloudcomputing-in-healthcare/
- Sivan R, Zukarnain ZA. Security and privacy in
cloud-based e-health system. Symmetry, 2021;23;13: 742.
- Current Use Cases for
Machine Learning in Healthcare, Microsoft Azure Blog, Microsoft, 2024;26.
- Introduction to BigQuery ML,
Google Cloud Documentation, Google, 2024.
- How Lumiata Democratizes AI
in Healthcare with Google Cloud, Google Cloud Blog, 2024.
- Transforming Healthcare in
the Cloud through Data Analytics Machine Learning, Cloud Blog, 11 Feb. 2019,
Accessed, 2024.
- Obermeyer Z, Emanuel EJ.
Predicting the future big data, machine learning and clinical medicine, New
England Journal of Medicine, 2016;375: 1216-1219.
- Kavakiotis I, Tsave O,
Salifoglou A, Maglaveras N, Vlahavas I, Chouvarda I, Machine learning and data
mining methods in diabetes research, Computational and structural biotechnology
journal, 2017;15: 104-116.
- Yu KH, Beam AL, Kohane IS. Artificial intelligence
in healthcare. Nature biomedical engineering, 2018;2: 719-731.
- Raghupathi W, Raghupathi V.
Big data analytics in healthcare: promise and potential, Health information
science and systems, 2014;2: 1.
- Healthcare Solutions
Playbook. Google Cloud, National Institutes of Health, Accessed, 2024.
- Borkowski AA, Wilson CP, Borkowski SA, Thomas LB,
Deland LA, Grewe SJ, Mastorides SM, Google Auto ML versus Apple Create ML for
histopathologic cancer diagnosis; which algorithms are better? 2019.
- Tigani J, Naidu S. Google
bigquery analytics, John Wiley & Sons, 2014.
- Create Machine
Learning Models in BigQuery ML, Google Cloud, 2024.
- Hosmer Jr DW, Lemeshow S.
Sturdivant RX. Applied logistic regression, John Wiley & Sons, 2013.
- Chen T, Guestrin C, Xgboost A. A scalable
tree boosting system, Proceedings of the 22nd acm sigkdd international
conference on knowledge discovery and data mining, 2016: 785-794.
- Sze V, Chen YH, Yang TJ, Emer JS. Efficient
processing of deep neural networks: A tutorial and survey, Proceedings of the
IEEE, 2017;105: 2295-2329.
- Teboul A. Health Indicators
Dataset, Kaggle, 2024.
- Powers DM. Evaluation: from precision, recall and
F-measure to ROC, informedness, markedness and correlation, 2020.
- Chicco D, Jurman G. The
advantages of the Matthews correlation coefficient (MCC) over F1 score and
accuracy in binary classification evaluation, BMC genomics, 2020;21: 1-3.
- Bustos K, Maazallahi A,
Salari M, Snir E, Norouzzadeh P, Rahmani B. Classifying and forecasting seismic
event characteristics using artificial intelligence, 2024.
- Adrian J, Mingxuan
Y, Ye T, Norouzzadeh P, Amini MH, Salari MA, Snir E, Rahmani B. COVID19 does
not correlate with the temperature, International Journal of Biomedical
Engineering and Science (IJBES), 2024;11.
- https://survey.stackoverflow.co/2024
- Wagholikar KB, MacLaughlin KL, Mandrekar J, Simha
VV, Liu H. Use of automatic SQL generation interface to enhance transparency
and validity of health-data analysis. Informatics
Med Unlocked, 2022;31: 100996.
- Tarbell R, Agrawal S, Xie Q, Sun L, Talukder A, Wallace BC, et al. Towards understanding the generalization of medical text-to-SQL models and datasets. AMIA Annu Symp Proc, 2023;2023: 669-678.
- https://cloud.google.com/bigquery/docs/auto-preprocessing