Begomovirus Recombination: Mechanisms Driving the Emergence of Novel Strains in Plant Pathogenesis
Abstract
Begomovirus, a genus of the Geminiviridae family, is one of the most important plant pathogens that spread to many economically significant crops. They are small, ssDNA viruses that harbor circular genomes and are mainly transmitted by whiteflies (Bemisia tabaci). Recombination, especially homologous and copy-choice recombination, is a key factor in the evolution of Begomoviruses. Recombination enables the virus to obtain a level of genetic diversity that leads to new strains that may have varying degree of pathogenicity. This process allows Begomoviruses to evade host resistance mechanisms, adapt to new host species, and rapidly respond environmental changes. The need to manage Begomovirus-encoded diseases is often made more difficult by the emergence of rapidly evolving recombinant strains of strain where recombination can potentially generate strains with increased virulence, a broader spectrum of hosts or strains that are resistant to commonly used control measures such as resist ant crop varieties or chemical treatments. Recombination is a major factor influencing the evolution of Begomoviruses and has an important role in plant disease management. This review highlights the importance of recombination in Begomovirus evolution, its implications on plant disease management It identifies increasingly biotechnological modes of eradicating the deleterious effects of viral recombination. Molecular diagnostic tools such as PCR and sequencing deliver a means for early recognition of recombinant strains. Genetic editing technologies like CRISPR/Cas9 will enable development of resistant crops and offer potential for restricting viral recombination. RNA interference (RNAi) is another example of a method used in silencing viral genes and replication. Transgenic crops also have potential for durable resistance against recombinant Begomovirus strains. This knowledge is vital to producing viable integrated virus control initiatives and ensuring a continued food supply worldwide by protecting crops from viral diseases.
Keywords: Begomovirus, recombination, novel strains, plant pathogenesis, biotechnology, viral evolution.
1. Introduction
Begomoviruses
are members of the family Geminiviridae, a large group of plant viruses that
cause serious diseases to many different dicots and other agricultural crop
species1-3. The genomic
characterization of these viruses is their small circular single stranded DNA
genomes4. Begomoviruses are one
of the most ubiquitous viruses affecting crops in the tropics and subtropics,
with high levels of transmission by whiteflies (Bemisia tabaci) and
productivity losses seen on economically important crops including but not
limited to; cotton, tomatoes, peppers, beans and ornamental plants5. Begomoviruses pose a serious threat to
global agriculture because these infections can lead to severe yield losses6. Begomovirus encompass highly diverse
viral species, with all sorts of bipartite and monopartite genomes (depending
on the species). Bipartite Begomoviruses have a dual component of DNA, whereas
monopartite viruses consist of one single genome, rendering them interesting
objects of study to explore their replication, transmission and pathogenesis7. These viruses tend to have the greatest
economic impact in areas that are dependent upon the agriculture of infected
crops, as they significantly decrease crop productivity and increase the
expenditure related to disease management8.
In (Table 1) major Key Begomovirus associated diseases, host range and
vectors, symptoms and economic damages.
Table 1: Major Begomovirus diseases, hosts, vectors, symptoms and
economic impact.
|
Virus/Disease |
Host Crops |
Vector |
Key Symptoms |
Economic Impact |
References |
|
Cotton Leaf Curl Kokhran Virus
(CLCuKoV) |
Cotton, okra, chili, cucumber,
papaya, soybean, luffa, and ornamentals Plants |
Whiteflies (Bemisia tabaci) |
Leaf curling, vein thickening,
enations |
Up to 100% yield loss |
9,10 |
|
Cotton Leaf Curl Multan Virus
(CLCuMuV) |
Cotton, okra, chili, cucumber,
papaya, soybean, luffa, and ornamentals |
Whiteflies |
Leaf curl, vein thickening,
stunting |
100% crop losses in South Asia
mostly |
9,11,12 |
|
Tomato Leaf Curl New Delhi Virus
(ToLCNDV) |
Tomato, cucurbits, Cotton |
Whiteflies |
Leaf curling, stunting, reduced
fruit set |
50% yield reduction |
13,14 |
|
Tomato Yellow Leaf Curl Virus
(TYLCV) |
Tomato, pepper, eggplant, potato,
and weeds like datura |
Whiteflies |
Yellowing, leaf curl, stunted
growth |
Major global tomato losses |
14,15 |
|
Cassava Mosaic virus (CMV) |
Cassava, Cotton |
Whiteflies |
Mosaic, distortion, stunted growth |
$1 billion annual losses |
8,16 |
|
Okra Enation Leaf Curl Virus
(OELCuV) |
Okra, Cotton |
Whiteflies |
Leaf curl, enations, reduced yield |
Significant yield losses in
Asia/Africa |
5,17 |
|
Chilli Leaf Curl Virus (ChiLCV) |
Chilli, pepper, Cotton |
Whiteflies |
Leaf curl, puckering, stunting |
Major yield reduction |
5,18 |
Recombination is an important
evolutionary mechanism for various viruses including the Begomoviruses19. This is the mechanism by which two viral
strains of distinct genetic heritages swap genetic information during
infections within the same host plant. In turn, this leads to the formation of
chimeric viruses with genetic components from each parental strain20. Because it enables rapid generation of
genetic diversity, recombination contributes to viral adaptation in altered
environments18, as well across species
barriers (S3) and host resistance mechanisms21.
With Begomoviruses, recombination can result in the emergence of new strains
with modified pathogenicity and virulence, host range or resistance to control
methods18. Recombination is
important for the evolution of Begomoviruses because it allows those viruses to
rapidly adapt to new environmental conditions and therefore, becomes harder to
control with conventional methods22.
Understanding recombination is therefore critical to developing management strategies
for Begomovirus diseases, particularly in an era where food security and
climate change are top global concerns23.
However, despite the significance of
Begomovirus recombination, there are important gaps in knowledge. Although
recombination is known as an important mechanism for the evolution of
Begomoviruses, the molecular mechanisms involved in these new recombination
remains obscure24. The specific
genomic regions of Begomoviruses that are prone to recombination have not yet
been determined and the host environmental factors affecting recombination
remain largely unexplored13.
Further, how these recombination events drive the emergence of new pathogenic
strains is not well understood. Without this knowledge, forecasting new
Begomovirus strains that may lead to more devastating diseases or managing them
are impossible.
We will primarily focus on the
molecular basis of Begomovirus recombination and its contribution to the rapid
evolution of new viral genotypes. Here we review the most recent advances in
knowledge regarding Begomovirus recombination, with an emphasis on how these
viruses evolve and how recombination leads to emergence of new strains, often
having different pathogenic potential, as well as discussing applications from
this perspective in plant disease management and biotechnology. The molecular
basis of Begomovirus recombination has implications for not only basic virology
research, but also for applying the knowledge to deploy new biotechnological
applications and control strategies such as the use of transgenic plants, viral
resistance, and RNA interference technologies against Begomovirus infections.
The Review serves to fill in research gaps that are currently left unaddressed
while laying out a map for future studies looking at viral evolution and its
effects on agriculture and biotechnology.
2. Begomovirus Recombination Mechanisms
2.1. Molecular basis of recombination
Recombination is one of the most
important mechanisms through which Begomoviruses evolve, providing genetic
diversity that ultimately allows viruses to adapt in new environments. Understanding
the molecular basis of recombination in begomoviruses occurs during the
replication of viral genomes and when two different strains infect the same
host plant. This co-infection enables genetic transfer leading to recombinant
viruses that contain dissimilar genomes of both the parent strains. The
homologous recombination and copy-choice recombination are the two common types
of recombination seen in Begomoviruses25.
Homologous recombination, this
occurs when two viral genomes contain regions of sequence identity and are able
to align and exchange genetic information during replication. In most cases,
this occurs in areas with high sequence homology allowing for the recombination
machinery to identify and swap homologous segments of DNA26.
Copy-Choice
Recombination, the viral polymerase continues the synthesis of a new genome by
switching from one template to another in this case copying genetic material
from both an original template and a different strain of virus. Such
recombination can produce hybrid viral genomes that contain segments from both
parent strains27. This
contributes to the generation of genetically diverse populations of virus,
which enables the virus to adapt (Figure 1).
Figure 1: Major recombination mechanisms in begomoviruses (A) Homologous recombination (RecA-mediated) occurs through
sequence alignment between related parental genomes, followed by crossover and
strand exchange, resulting in recombinant genomes with mixed genetic segments.
(B) Copy-choice recombination (Rep-mediated) takes place during rolling-circle
DNA replication, where the viral replication-associated protein (Rep)
facilitates template switching between donor and acceptor DNA strands,
generating recombinant genomes.
Recombination is a very intricate
process that includes several kinetic factors: some of them are enzymes, others
are host proteins. Central to the recombination is the Begomovirus replicase,
which synthesizes new DNA copies of viral genome information. This enzyme can
also contribute to homologous recombination or copy-choice recombination,
depending upon replication dynamics and the template present28. Furthermore, host cellular proteins that
play a role in DNA repair and replication processes, such as DNA polymerases
and recombinase enzymes, also play a crucial role in repairing and
restructuring the viral genome during homologous recombination. These host
factors involved in replication, such as RNA polymerases and other regulatory
proteins might also modulate the recombination process by supplying the
conditions that allow for the exchange of genetic material29.
2.2. Role of recombination in viral evolution
Recombination is important in viral
evolution as it creates genetic diversity, especially within begomoviruses30. Such diversity gives the virus the
capacity to quickly evolve in response to selective pressures, including host
resistance mechanisms, environmental pressures or changes and new vectors. In
contrast to point mutations, recombination can trigger the quick emergence of
new viral variants that may have beneficial attributes, including higher
virulence, an expanded host range or escape from control measures31.
Recombination allows Begomoviruses,
on the other hand, to be more rapid tractor toward the new characters than
mutations ever could. Exchanging genetic material between strains also allows a
virus to gain genes that help it cope with host immunity or persist in new
ecological niches32. The fast
adaptability has made Begomoviruses a major threat to crops because new strains
with new pathogenicity appear in short time, and disease management becomes
very complicated18,20.
As an example, mutant strains of
Begomoviruses recombinants have been revealed that display transformed host
specificity and quality with the ability to contaminate resistant plant
species. Recombination, for example, can generate strains that are more virulent,
damaging crops and giving rise to more diseases that matter. With the evolution
of these recombinants, they could also escape resistance control, such as
resistant crops or chemical treatment leading to higher management complexity33.
2.3. Recombination hotspots
Although recombination does occur
throughout the genomes of Begomovirus, some regions are more predisposed to
such events due to their genomic architecture and functionality. Recombination
hotspots are usually located in genome regions that are important for virus
replication and host interactions. An example of this region is the intergenic
region (IR) that is situated between both elements of the bipartite genome
organizations found in many Begomoviruses2,34.
The IR is common to replication and transcription of the virus and harbors
conserved features that enable recombination to occur among closely related
viral strains. Another recombination hotspot is the C4 gene, which encodes a
protein associated with pathogenesis and manipulation of host cellular
processes. Mutations or recombination on that gene may cause major alteration
of the virus possible to reach a number of plant's species35.
These recombination hotspots are
important because they signify the parts of the genome that are most engaged in
the virus’s infection and proliferation ability within host plants. Such
recombination can result in the generation of viral strains with modified
pathogenicity, potentially allowing for a wider range of plant species to be
infected and/or their virulence. A deep understanding of these hotspots is
needed to predict evolutionary behavior of Begomovirus strains, and to devise
counterstrategies for the impact of recombinant emerging strains17.
Homologous recombination and copy-choice
recombination, the molecular mechanism of Begomovirus recombination, play a key
role in the emergence of new viral types. The underlying mechanisms allow
Begomoviruses to create genetic variance, which allows new strains with
different pathogenicity to emerge. This knowledge of the distribution and
nature of hot spots for recombination across the viral genome can also assist
researchers in tracking how strategies have evolved with evolutionary
adaptation as well, providing new approaches to controlling risks posed by these
emerging Begomovirus strains36.
3. Emergence of Novel Strains
3.1. Mechanisms driving novel strain
emergence
Genetic material can be exchanged during
co-infection by two different strains of the virus within one host, resulting
in new viral variants. It is the reason that more virulent strains arise by
bringing across useful characteristics from both parent strains during this
recombination process. The new strains may better evade plant defense or have
greater adaptability to environmental changes, making them more fit in wild
populations37. Recombination has
important consequence, which is that new strains escape host resistance. Over
the course of time, host plants gain resistance either naturally or through
breeding. But in recombinant viruses, these resistance mechanisms are overcome
because the new genetic traits allow it to infect species that were previously
resistant. Strains that evolve through recombination could also change their
replication machinery or coat protein structure to decrease susceptibility to
plant immune responses (Figure 2). This allows the virus to infect pl.
species across from a wider spectrum of host plant or escaping pre-existing
R-genes that were previously effective against earlier strains38.
Figure 2: Recombination-driven
emergence of novel begomovirus strains. It shows how co-infection of a host
plant by different viral strains can lead to recombination during replication,
generating recombinant genomes and resulting in novel strains with altered traits
such as increased virulence, expanded host range, overcoming host resistance,
and resistance to control measures.
In
addition, recombination can have a substantial impact on virus-host
interactions and enable Begomoviruses to alter their interaction with plant
defence systems. In this way, recombined strains may adopt new strategies to
hijack host cellular machinery and ultimately increase their ability to
replicate and spread within the plant. This results important changes in the
plant and could lead to a more aggressive version of disease, as pathogen
utilizes resources from host plant at higher rates also influencing infection
levels and symptoms severity39.
3.2.
Examples of novel begomovirus strains
A
notable instance of Begomovirus recombination giving rise to new, more virulent
strains is the Cotton leaf curl Multan virus (CLCuMV). Recombination among
different CLCuMV strains has been associated with higher virulence and wider
host ranges. At the beginning, CLCuMV was only to cause infection in cotton but
via recombination the virus has started remixing and being able to infect a
range of other crop species including tomato, pepper and several vegetable
species. As a result of this recombination event, cotton and many other crops
have been subject to major losses due to the economic importance of
Begomoviruses11.
Apart
from cotton, these novel strains of CLCuMV have now spread to a number of crops
in several regions such as Pakistan, India, China and also the Philippines and
Thailand. In these areas, the virus has evolved to infect other plant species
which adds a layer of difficulty for management of the two diseases. Recombination
events increasing the ability of CLCuMV to infect a wider range of hosts are
one of the best examples demonstrating how recombination produces strains with
enhanced virulence and adaptability, which results in an increased threat to
agriculture. This poses important economic significance, especially for cotton
producing countries where cotton is a very important cash crop, as well as
areas with large acreage of tomatoes and peppers12.
Likewise,
recombination among other Begomovirus species have produced strains with
changed pathogenicity. One notable case is the recombination of Tomato yellow
leaf curl virus (TYLCV) strains followed by emergence of new virulent viral
variants that infect crops in previously TYLCV-uninfected areas. The emergence
of such new strains renders existing control measures ineffective in managing
agricultural Begomovirus diseases, thereby complicating its management. The
emergence and dissemination of recombinant strains pose considerable challenges
to the global agriculture community in controlling viral infections
effectively, underscoring an urgent need for novel strategies for viral
management and surveillance15.
3.3. Geographic distribution of novel
strains
Recombinant Begomovirus strains have
spread globally and have been associated with severe outbreaks in areas where
susceptible crops are grown40.
This proliferation of new strains is evident in South Asia where the
identification of new Begomovirus genotypes due to either strain recombination
or movement of whitefly vectors, has been reported41,5
specifically from Pakistan and India. More virulent strains such as those
causing Cotton leaf curl disease; have spread rapidly in these regions. High
density of host plants, multiple infections of diverse viral strains and
appropriate conditions that favor whitefly dispersal are responsible for the
increased proportion of recombinant strains42.
Private sector-mediated global trade and
agricultural practices, including the movement of infected plant material as
well as extensive production of susceptible crops, further facilitate the
transboundary movement of such novel strains43.
The migration of the whitefly vectors has been increased due to climate changes
and near agricultural area17. The
understanding of factors influencing the geographical distribution of newly
dispersed strains is critical to limit the spread and manage their detrimental
effects on global agriculture44.
Recombination, which leads to the
emergence of new Begomovirus strains, mainly allows viruses to escape host
resistance and adapt to new environments, thereby increasing virulence45. The dynamic nature of the Begomovirus
evolution is reflected in the case of Cotton leaf curl Multan virus and other
recombinant strains. An added concern is that the widespread geographic
distribution of the new strains makes the case for a global surveillance and
co-ordinated control strategy for plant disease to be put in place first, if
the threat they pose to agricultural production and food security is to be
mitigated.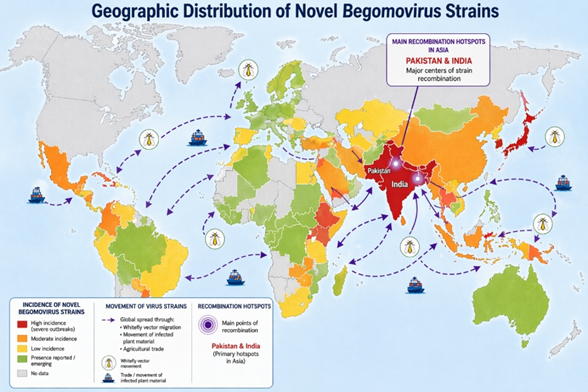
Figure 3: Global
geographic distribution and movement of recombinant Begomovirus strains.
4. Implications for Plant Pathogenesis
4.1.
Disease dynamics
The
impact of recombination on plant disease dynamics is massive as new Begomovirus
strains occur through this mechanism. Recombinant strains, which consist of a
mixture of genetic material from multiple viral strains, more virulent, causing
more severe disease manifestations and rapid progression. Recombination may
allow Begomoviruses to circumvent host immune defenses, particularly in the
case of new viral variants that escape resistance mechanisms acquired by
selective breeding30 or those acquired through natural plant defense systems38. Specifically, recombination can alter
coat proteins or machinery involved with virus replication; these mutations
allow the virus to infect new host species, or evade plant defense responses10. This dynamic responsiveness to
environmental and host alterations is crucial in the propagation and
manifestation of viral diseases, particularly when novel strains possess
increased infectivity and pathogenicity.
4.2.
Impact on crop yields
In
agriculture, recombinant Begomovirus strains and their economic impact are
significant. These strains act more aggressively and result in substantial crop
yield losses. Begomovirus-induced diseases like the ones occurred from Cotton
leaf curl Multan virus (CLCuMuV) which can severely adversely affect cotton by
reducing fiber quality and causing yield loss. This demonstrates the potential
for recombination of the virus to infect a broad variety of host plants, which
in turn represents an ongoing food security challenge46. The increased cost of disease management
is attributed to the emergence of strain diversity through recombination that
enables the virus to adapt to new conditions or evade resistance. Farmers may
therefore have to undertake more frequent pesticide application, purchase of
genetically resistant crops or changing the crop management practices, each of
which adds up to greater costs. This situation is aggravated by the diffusion
of these recombinant strains via trade and agricultural practices across
regions around the world, which translates into economic losses globally47.
4.3.
Host-pathogen co-evolution
Recombination
influences not just the virus but also provokes host-pathogen co-evolution.
Plants also have constantly adapted their defense systems to resist virulent
infections, and viruses, mainly Begomoviruses evolve quickly in order to overcome
those defenses48. This promotes
co-evolution because recombination allows for viral evolution of new immune
evasive strategies. For instance, Begomoviruses can change the conformation of
their coat proteins or adjust their replication mechanisms to evade host defense49. This might prompt host plants to evolve
new resistance genes to defend against new viral strains such as BSMV-F.
Similar to other evolutionary arms races, this conflict between the virus and
its host ultimately shapes the potential persistence of Begomoviruses in
agricultural systems. Due to the emergence of new strains, plant breeders are
challenged to develop more sophisticated resistance strategies49, which makes the management of viral
diseases even more complex.
Recombinants
between Begomovirus strains cause serious threats to plant pathogenesis. Agriculture
and Disease management In addition to the improved virus propagation capacity,
these strains also provided additional challenges in managing viral diseases.
It has widespread economic implications, imposing higher costs on farmers and
greater risks to yields16.
Additionally, the co-evolutionary dynamics between Begomoviruses and their host
plants indicate that without proper management approaches this cycle of virus
adaptation and resistance in host will continue, possibly resulting in more
virulent strains with an ever-increasing epidemic potential that can lead to
polycyclic diseases. Most importantly, insights into the mechanisms governing
recombination will better inform virus-host interactions that are necessary for
sustainable management of Begomovirus diseases in future.
5. Biotechnology Approaches to Understanding and Controlling
Begomovirus Recombination
5.1. Diagnostic tools
Molecular diagnostics are critical in
the detection of recombinant Begomovirus strains that enable early diagnosis
and management of viral diseases. Recombinant strains are uniquely fielded
using techniques such as polymerase chain reaction (PCR) and next-generation
sequencing (NGS) statistics50.
PCR-based approaches can certainly confirm the presence of Begomovirus DNA in
plant tissues, whilst sequencing enables detailed analysis of viral genomes
that allows identification of recombination events and on-going emergence of
novel strains. Such PCR assays that can target specific genomic regions
susceptible to recombinant such as the intergenic region (IR) and the C4 gene
have been used successfully to differentiate recombinant from parental strains51. Furthermore, sequencing techniques allow
us to capture the entire scenes of recombination hotspots on viral genomes and
provide insights into how genetic transmissions happen and what regions are
more susceptible to such events52.
Figure 4: Workflow for detection and characterization of begomovirus recombinants
using molecular diagnostics and bioinformatic analysis.
5.2.
CRISPR/Cas9 and gene editing
The
development of new gene editing technologies, especially CRISPR/Cas9 (clustered
regularly interspaced short palindromic repeats/CRISPR-associated protein 9)
have opened exciting possibilities for controlling the high frequency of
Begomovirus recombination and improving plant resistance against these
genotypes. CRISPR/Cas9 systems may be used for targeted cleavage of a site in
the Begomovirus genome, which could derail the recombination events that confer
virulence. Appropriate editing of viral genomes could also restrict the ability
to undergo recombination, which in turn could limit the emergence of novel
pathogenic strains53. In
addition, CRISPR/Cas9 can be applied to plants by modifying plant genes behind
the host-pathogen relationship, avoiding infection. For instance, editing plant
genes encoding susceptibility factors54
or increasing the expression of resistance-activating genes55 would endow crops with capabilities to
resist viral infections driven by recombination events.
5.3.
Viral resistance through transgenics
One
of the most relevant biotechnological methods for curtailing the emergence of
new recombinant Begomovirus strains is through the use of transgenic crops55. Introduction of particular genes into
plant genomes has, to some extent, produced Begomovirus-resistant
genetically-modified crops. For one, the C4 gene from Tomato yellow leaf curl
virus (TYLCV) has been introduced into cotton and other crop plants which gave
them resistance against Begomovirus-related diseases. This would mean that
transgenic plants are either preventing the viral replication cycle or they are
increasing the plant's immune reaction, limiting the efficiency of infection
and spread by Begomovirus. An important tool in the management of these
ever-changing viruses is the use of transgenic crops tolerance to recombinant
Begomovirus strains14.
5.4.
RNA interference
An
additional strong biotechnology-based tool against Begomovirus recombination is
RNA interference (RNAi) that could help to minimize its effects on crop health56. RNAi silences selected genes of a virus
to stop it multiplying in a host plant. This method consists of a form of RNAs,
double-stranded RNA (dsRNA) that is introduced into the plant and these strands
activate and stimulate the plant RNAi tools to degrade viral RNA. RNAi can
reduce viral load by silencing essential genes for replication, such as those
related to the replicase or C4 gene57.
Moreover, RNAi can silence specific recombination-prone zones in the
Begomovirus genome, preventing virus emergence of new more virulent strains58.
5.5.
Synthetic biology approaches
Synthetic
biology is a new technology that integrates aspects of engineering, genomics
and biochemistry to construct biological systems in novel ways59. For Begomovirus recombination, a
synthetic biology approach could allow for the creation of new combinations of
viral genetics either coded into modified viral systems or be constructed as
engineered genomes together with designed built-in controls to prevent any
further potential recombination events. As an example, this could result in the
synthesis of engineered viral systems that contain stable and non-recombinant
genomes such that, within a controlled system, evolution into a new strain is avoided60. Synthetic biology could also be used to
design crops with boosted immune responses or even synthetic viral “decoys”
that lure and destroy the virus, preventing infection of healthy crops. These
strategies could be powerful not only for regulating viral evolution, but also
establishing robust agricultural ecosystems and safeguarding against new viral
challenges61.
Intriguingly,
there are a number of interesting and potentially useful biotechnology tools
which could aid in elucidating the recombination occurrence and/or control this
organism. These approaches, ranging from molecular diagnostics and CRISPR/Cas9
gene editing to RNAi and synthetic biology, are paving the way for novel
strategies that will allow us to tackle viral evolution and enhance plant
resistance. Our findings provide a foundation for future work and important
biotechnological interventions to show their value in the emergence of
Begomovirus that evolve through recombination (Table 2), and will be
critical for maintaining the sustainability of global agriculture while
reducing economic losses caused by viral diseases62.
Table 2: Biotechnological
approaches for detection and control of Begomoviruses.
|
Approach |
Principle |
Application |
Advantages |
Limitations |
References |
|
PCR
/ qPCR |
DNA
amplification |
Detection
and quantification |
High
sensitivity |
Lab
setup required |
50,63 |
|
NGS |
Genome
sequencing |
Recombination
detection |
High
resolution |
Expensive |
51,53 |
|
CRISPR/Cas9 |
Genome
editing |
Virus-resistant
crops |
Precise |
Regulatory
issues |
53,54,64 |
|
RNAi |
Gene
silencing |
Virus
suppression |
Specific |
Variable
efficiency |
56,57 |
|
Transgenic
crops |
Gene
insertion |
Resistance
development |
Long-term |
Regulatory
barriers |
55,65 |
|
Synthetic
biology |
Engineered
systems |
Virus
control |
Innovative |
Early-stage
tech |
59,61 |
6. Challenges in Begomovirus Management and Future Directions
6.1.
Recombination control challenges
Frequent
recombination is a feature of Begomoviruses rendering that difficult to
control, especially in farming systems. Recombination facilitates rapid
adaptation of Begomoviruses to new hosts, environments and resistance
mechanisms, it is nearly impossible to predict the emergence of novel strains.
In contrast to the more gradual mutations, recombination may give rise to
strains with increased virulence or host adaptation all at once66. This is particularly problematic for the
management of disease because recombination can break down durable resistance
in crops67, and thus requires
ongoing new control strategies. The ability of Begomoviruses to rapidly evolve
makes traditional approaches such as the development of resistant cultivars or
chemical control ineffective upon the emergence of a new recombinant strain67. This means that successful manipulation
of recombination in Begomoviruses relies on knowledge of the molecular basis of
this process but also, at the same time, new adaptive and flexible strategies
to control it based on rapidly evolving targets.
6.2. Barriers to biotech solutions
Though biotechnology looks promising in
terms of tools for the management of Begomovirus, several hurdles limit its
scope. These still poses a major barrier due to regulatory hurdles, especially
in the domain of GMOs and transgenic crop approval. The legislative buffers in
different countries that oversee GMOs differ, with the strict testing and
safety assessments that are required delaying resistant crops65. In addition, there are worries about the
environmental consequences of the introduction of genetically modified crops
into ecosystems, especially unintentional effects on biodiversity or that wild
virus populations can develop resistance. There are also persistent technical
hurdles, especially for site-specific targeting of viral genes or stable
genetic engineering of resistance traits across the target crop genomes. And,
despite CRISPR/Cas9 and other advanced tools, the attainment of effective,
stable and large-scale expression of resistance in crops remains an enormous
challenge that will require substantial research and development. Such barriers
highlight the relevance of work forums to ensure cooperation between
researchers, policy-makers and agriculture stakeholders in order to overcome
hurdles and facilitate the implementation of biotech solutions68.
6.3. Future research directions
Several gaps remain that have hitherto
inhibited the effective management of Begomovirus', even given recent advances
in our understanding of this group of viruses. Better tools for diagnosis are
essential to identify recombinant strains in early stages, especially in areas
where Begomovirus is endemic69.
More sensitive and rapid diagnostic techniques would enable enhanced
surveillance and significantly quicker responses to emerging strains.
Resistance breeding is another component that need further studies70. Although transgenic approaches hold
promise, the need for durable resistance against Begomoviruses still relies on
conventional breeding for novel sources of resistance. A focus on new methods
for breeding crops with novel germplasm to bring these genetic traits into
commercially cultivable plants will be beneficial, and many knowledge gaps
exist regarding how resistance can be obtained from wild relatives of crop
species71. Also, how recombinant
viruses are characterized is an area still under investigation.
Greater‑resolution genome mapping of
begomovirus strains, especially recombinants, will help localize potential
recombinogenic hotspots and re‑assess the role of these regions in the evolution
of novel virulent variants72.
Future research should also investigate vector control strategies, as
whiteflies are the primary vectors of begomoviruses and play a central role in
the spread of these diseases. Information on whitefly population dynamics and
their interactions with begomoviruses is essential to understand how to
effectively limit the spread of the virus across different geographic regions.
Despite the potential of biotechnology
to/for control Begomovirus diseases, regulation, environmental concerns and
technical aspects will require further multidisciplinary effort to be overcome.
Advancing diagnostic techniques, resistance breeding and the characterization
of recombinant viruses will be key to designing more effective control
strategies for these rapidly diversifying plant pathogens.
7. Conclusion
7.1.
Summary of key insights
Recombination
underpins Begomovirus evolution through its ability to rapidly generate
molecular diversity2,19. This
process is responsible for the generation of new viral strains that can bypass
host resistance, be incorporated into new climatic and environmental
conditions, and become impervious to control strategies. Since the majority of
Begomoviruses are bipartite or multipartite, recombination enables faster
evolution than mutations alone and promotes viral property changes to be
disseminated rapidly. Consequently, the role of Begomovirus evolution by
recombination can have a profound impact on plant virology73, making it more difficult for effective
disease management and increasing the threat to world agriculture. The rapid
evolution and adaptation potential of Begomoviruses highlights the importance
of AV analysis to aid continuous monitoring, deployment of new management
strategies and development of agri-policies that consider viral diversity and
recombination74.
7.2.
Biotechnological promise
Biotechnology
could have an important role in the reduction of adverse effects of viral
recombination on Begomovirus disease control. Molecular diagnostics, gene
editing using CRISPR/Cas9, RNA interference (RNAi), and the development of
transgenic crops are some major biotechnological tools that can be used against
Begomovirus75. CRISPR/Cas9 is for
altering viral resistance genes; RNAi technology silences any virus genes
giving rise to recombinant strains. In addition to the health benefits,
genetically modified crops that are intrinsically resistant to specific strains
of Begomovirus would also minimize the economic impact of these viral diseases.
These biotechnological solutions remain subjected to regulatory and technical
hurdles, but they could provide fundamentally new approaches to the management
of Begomovirus infections by providing tools that can slow viral evolution and
their agronomic impact64,76.
7.3.
Final thoughts
Management
of Begomovirus strains, therefore, must adopt a multi-disciplinary approach in
combining virology, plant breeding and biotechnology with pest management
practices. This suggests that these viruses are facing substantial evolutionary
pressures via recombination; therefore, scientists need to adopt flexible and
adaptive approaches. This will include the generation of new diagnostic tool
and resistance breeding technologies, as well as employing novel
biotechnologies to limit spread recombinant strains and develop durable
resilient crops. International collaboration and cross-border knowledge sharing
will be fundamental to addressing the challenges posed by Begomovirus
infections, given their complexity, the global scale of their impacts, and need
for sustainable solutions towards this issue. Begomovirus management of the
future will rely on a more integrated method that uses advances in science and
agricultural practices to protect crops worldwide, keeping food secure.
8. Declarations
8.1.
Ethics approval and consent to participate
N/A
8.2.
Consent for publication
All
authors are agreed for publication of the article.
8.3.
Availability of data and materials
All
data is available and provided upon request to corresponding author.
8.5.
Conflicts of interest
There
is no conflict of interest.
8.6.
Funding
N/A
8.7.
Authors' contributions
FA
ER, MYM and MS idea and wrote this manuscript. MBA, EY, AM, MA review and
format the manuscript.
8.8.
Acknowledgements
All
authors are very thankful to Virology lab members for critical review.
9. References
2. Malik MY. Molecular Diagnostics
and Management Strategies for Begomovirus-Induced Cotton Leaf Curl Disease: The
Role of Weeds as Reservoirs and Vectors. Biol Times, 2026;5(1): 4-6.
3.
Yasmeen E, Ghaffar B, Bibi Z, et
al. Genomic Plasticity and Molecular Evolution of Begomoviruses. Biol Times,
2026;5(2): 21-23.
10.
Jan A, Razzaq F, Umair M, et al.
Cotton leaf curl disease: Pathogen diversity, whitefly ecology, and integrated
management approaches. Planta Animalia, 2025;4(4): 363-371.
13.
Behera BP, Mallick K, Behera A,
et al. Plant Viruses in a Changing Climate: Tomato Leaf Curl New Delhi Virus
Dynamics, Evolution, and the Myth and Reality of Beneficial Interaction. In:
Tomato Leaf Curl New Delhi Virus (ToLCNDV) Insights into Virome Dynamics and
Management. Singapore: Springer Nature, 2025: 231-253.
35.
Krishnan N, Kumar RV, Karmakar P,
et al. Co infection and recombination driven emergence of begomovirus complex
causing mosaic disease in muskmelon (Cucumis melo L.). Mol Biol Rep,
2026;53(1): 445.
40.
Reddy MK, Nagendran K, Kumari S,
et al. Global scenario of begomovirus diseases in vegetable crops. Veg Sci,
2024;51: 43-53.
48.
Sanjuán R, Domingo Calap P.
Mechanisms of viral mutation. Cell Mol Life Sci 2016;73(23):4433-4448.
49.
Munshi A. Geminiviruses:
Exploring the Evolution Host Genetic Elements and Host Virus Interactions.
Raleigh, NC: North Carolina State University, 2022.
56.
Sharma R, Das T, Devi K, et al.
RNA Interference (RNAi) and Other Molecular Approaches for Tomato Leaf Curl New
Delhi Virus Control. In: Tomato Leaf Curl New Delhi Virus (ToLCNDV) Insights
into Virome Dynamics and Management. Singapore: Springer Nature, 2025: 255-270.
60.
Zaidi
SSEA, Mansoor S. Viral vectors for plant genome engineering. Front Plant Sci,
2017;8: 539.
66.
Fortes IM, Díaz Martínez L,
Moriones E, et al. Virus-Host Interactions and Genetic Exchange in Mixed
Infections of Tomato Yellow Leaf Curl Virus (TYLCV), Tomato Leaf Curl New Delhi
Virus (ToLCNDV), and Tomato Chlorosis Virus (ToCV). Agronomy, 2025;15(5): 1006.
68.
Raza A. Genetically Modified
Crops: Benefits, Risks, and Regulatory Perspectives. Front Agric, 2024;1(2):
252-282.
69.
Bharathi L, Biju TS, Viswanathan
A. Diagnostic strategies in begomovirus detection from traditional assays to
modern technologies. Heliyon, 2026;12(6).
75.
Barupal M. Contemporary
biotechnological interventions in plant disease management. J Mycol Pl Pathol,
2025;55(2): 103-120.